Diversidad genética de Oenothera drummondii (Onagraceae), una herbácea de dunas costeras: implicaciones ecológicas y evolutivas
Raquel Aurora Hernández-Espinosa a, Jorge González-Astorga a, *,
Yessica Rico b, Juan B. Gallego-Fernández c
a Instituto de Ecología A.C., Red de Biología Evolutiva, Laboratorio de Genética de Poblaciones, Carretera Antigua a Coatepec No. 351, El Haya, 91073 Xalapa, Veracruz, Mexico
b Instituto de Ecología A.C., Centro Regional del Bajío, Red de Diversidad Biológica del Occidente Mexicano, Av. Lázaro Cárdenas No. 253, 61600 Centro, Pátzcuaro, Michoacán, Mexico
c Universidad de Sevilla, Departamento de Biología Vegetal y Ecología, Av. de la Reina Mercedes, 6, 41012 Sevilla, Spain
*Corresponding author: jorge.gonzalez@inecol.mx (J. González-Astorga)
Received: 09 June 2025; accepted: 31 October 2025
Abstract
We studied the diversity and genetic structure of Oenothera drummondii Hook. (Onagraceae), a dune plant with a mixed reproductive system, across 9 populations using 10 microsatellite markers. Plant genetic diversity is governed by intrinsic factors (i.e., reproductive system and dispersal) and extrinsic factors (i.e., population fluctuations and founder effects). We found moderate to low genetic diversity, with southern populations showing lower diversity, and northern populations higher. A separate study on self-compatibility revealed higher selfing in the south and lower in the north, suggesting a latitudinal gradient that may reduce genetic diversity in southern populations. Peripheral populations showed reduced diversity and greater differentiation, likely due to increased isolation and limited gene flow. Central populations near the Texas origin exhibited the highest diversity. Populations from Baja California (O. drummondii subsp. thalassaphila) formed a genetically distinct group, suggesting a separate species. Overall, genetic patterns in O. drummondii genetic diversity reflect historical and ecological influences, including mating system variation, floral traits, and pollinator dynamics. These findings support hypotheses such as center-periphery dynamics and climate-driven historical processes (e.g., post-glacial expansion), which may shape the species’ genetic landscape and suggest possible local adaptations to environmental changes.
Keywords: Neutral genetic diversity; Coastal dunes; Microsatellites; Genetic bottleneck; Latitude; Selfing
Resumen
Evaluamos la diversidad y estructura genética de Oenothera drummondii Hook. (Onagraceae), planta de dunas costeras con sistema reproductivo mixto; se usaron 9 poblaciones y 10 marcadores microsatélites. La diversidad genética en plantas está influenciada por factores intrínsecos (dispersión y sistema reproductivo) y factores extrínsecos (fluctuaciones poblacionales y efecto fundador). Encontramos diversidad genética moderada a baja, con menor diversidad en poblaciones del sur (Ojoshal) y mayor en el norte (Bolívar). Un estudio complementario mostró mayor autocompatibilidad en Ojoshal y menor en Bolívar, sugiriendo un gradiente latitudinal con incremento de autocompatibilidad hacia el sur, con posible reducción de diversidad genética. Las poblaciones periféricas presentaron menor diversidad y mayor diferenciación, asociada a menor flujo génico y mayor aislamiento; las poblaciones centrales, cercanas al origen de la especie en Texas, mostraron mayor diversidad. Las poblaciones de Baja California (O. drummondii subsp. thalassaphila) formaron un grupo genético distinto, que podría representar a una especie separada. En conjunto, los patrones genéticos de O. drummondii reflejan procesos históricos y ecológicos relacionados con el sistema reproductivo, características florales y las dinámicas de polinización, lo cual podría significar adaptaciones locales a cambios ambientales, y patrones de distribución genética asociados con ellos, como la hipótesis de centro-periferia.
Palabras clave: Diversidad genética neutral; Dunas costeras; Microsatélites; Cuellos de botella; Latitud; Autocompatibilidad
Introduction
Genetic diversity may be considered a crucial factor in determining the ability of populations to adapt and evolve, thus increasing their evolutionary potential (He et al., 2024). Understanding the drivers of genetic variation among plant populations is essential in evolutionary biology as this diversity is the foundation for adaptive potential and long-term survival (Chung et al., 2023; Wright, 1969). In plants, the factors that shape genetic variation can be divided into: a) intrinsic biological properties (e.g., the genetic recombination system, including ploidy level, reproductive system, and meiotic behavior; the mode of dispersal and pollination; and the life form), and b) extrinsic dynamic processes (e.g., fluctuations in population size due to bottlenecks, founder effect, invasions, and changes caused by ecological succession) (Duminil et al., 2009). The reproductive system is the intrinsic biological property suggested to be the major driver of genetic diversity in plants (i.e., heterozygosity and population genetic differentiation) (Koelling et al., 2011; Wright, 1969), as it influences the mating patterns within a population by determining the extent to which selfing can occur (Charlesworth, 2006; Raduski et al., 2012). Self-incompatible populations, for instance, are composed of outcrossing plants with a predominantly outcrossing mating system. Conversely, in partially or completely self-compatible plants, the mating system can span from outcrossing to mixed mating to complete selfing (Holsinger, 1991).
Small, low-density populations experiencing habitat fragmentation and isolation, with limited mate or pollinator availability, exhibit increased propensity for self-fertilization (Devaux et al., 2014; Whitehead et al., 2018). In situations where self-fertilization is evolutionarily advantageous, selection will favor changes in traits that facilitate selfing (e.g., reductions in flower size, reduced nectar, pollen and reduced herkogamy) (Opedal, 2018; Shimizu & Tsuchimatsu, 2015; Sicard & Lenhard, 2011). These changes can have consequences at the genetic level, including elevated rates of inbreeding, a reduction in genetic diversity within populations, and increased genetic differentiation among populations (Barrett & Harder, 1996; Ingvarsson, 2002). Furthermore, a principal consequence of predominant self-fertilization is the reduction in heterozygosity and intra-population genetic diversity, coupled with an increase in inter-population differentiation, when compared to self-incompatible plants (Hamrick & Godt, 1996). In contrast, clonal species are expected to exhibit higher levels of heterozygosity than self-incompatible species, but lower levels of polymorphism within the population and higher levels of differentiation between populations (Levin, 2012).
In addition to the intrinsic biological properties of the species, spatial distribution and demography strongly influence processes such as genetic drift, gene flow, and natural selection, which in turn shape the genetic characteristics of populations (Eckert et al., 2008). For example, species range size tends to influence gene flow and genetic differentiation between populations. Large distances between populations with large ranges would be barriers to gene flow (Lawrence & Fraser, 2020). Peripheral populations are predicted to exhibit reduced gene flow, greater isolation, and higher genetic differentiation. In contrast, populations in the center of the distribution would have higher gene flow and be less differentiated (Lawrence & Fraser, 2020). This may be exacerbated if the peripheral populations experience rapid cycles of colonization and extinction, and have associated bottleneck events or founder effects (Eckert et al., 2008). Dispersal ability also influences the amount of gene flow between core and peripheral populations. This has implications for the maintenance of genetic diversity. For example, species with large ranges but limited mobility have reduced gene flow between populations, resulting in greater genetic differentiation (Pelletier & Carstens, 2018).
The genus Oenothera (Onagraceae) is widespread with the majority of species concentrated in western North America (Overson et al., 2023), with some taxa extending to Central Mexico and South America (Wagner et al., 2007). Like other Onagraceae members, Oenothera species originated in the Nearctic region. Their diversification began 20 million years ago during the Miocene, an epoch characterized by colder and drier than the present. These conditions triggered both altitudinal and latitudinal forest retreats, creating ecological opportunities for herbaceous species that prefer open environments and can resist low temperatures and low humidity (Dietrich & Wagner, 1988).
The species of Oenothera contributed significantly to the early development of plant genetics, cytogenetics, and evolutionary biology. Since the work of De Vries in 1900 (Cleland, 1972), a great deal of information has been collected on the ecology, morphology, cytology, and genetics of the genus, giving it a great advantage as a model for study (Greiner & Köhl, 2014), especially on reproductive systems (Johnson et al., 2009; Raven, 1979; Wagner et al., 2007). The intricate evolutionary history of the genus reports several transitions in reproductive systems, which are diverse (Cleland, 1972; Johnson et al., 2011; Rauwolf et al., 2008). In addition, Oenothera has a unique genetic recombination system (Cleland, 1972). In some species, meiosis is accompanied by a rearrangement of chromosome arms that form structures known as rings. These rings segregate into the next generation and have significant implications for inheritance (Golczyk et al., 2014). They act as barriers to homologous recombination and alterations to the linkage balance (Overson et al., 2023; Rauwolf et al., 2008).
Our study focuses on Oenothera drummondii Hook. (Onagraceae), an herbaceous, short-lived perennial species belonging to the subsection Raimmania within Oenothera (Overson et al., 2023). The speciespresents different chromosomal configurations during meiosis, including the formation of bivalents and rings, but is not a permanent translocation heterozygote; so its reproduction is not expected to be clonal (Dietrich & Wagner, 1988). In contrast, the species displays a mixed mating system, encompassing both self- and cross-pollination (Dietrich & Wagner, 1988; Gallego-Fernández & García-Franco, 2021b). Despite exhibiting self-compatibility, its herkogamous large yellow flowers, and elevated stigma suggest predominantly outcrossing, particularly in North American populations (Gregory, 1964). Nevertheless, the species is capable of self-pollination in the absence of pollinators (i.e., sphingid moths and some hymenopterans) or under specific environmental conditions, such as wind, sand burial, and wave action during tropical storms, which are common in dunes and have an impact on the survival of individuals (Dietrich & Wagner, 1988; Gallego-Fernández & García-Franco, 2021; Gregory, 1964). Recent studies in North America showed that levels of self-compatibility vary from population to population, decreasing with increasing latitude (Gallego-Fernández & García-Franco, 2021b). Also, flower size decreased significantly with increasing latitude (Gallego-Fernández & García-Franco, 2021a). Given that all these characteristics could be reflected in the genetic diversity and genetic structure of the populations, we combined existing reports on life history traits, germination and self-compatibility levels, floral traits, dispersal modes, and genetic recombination system with our results of the genetic diversity and structure of 9 populations covering the North American distribution of O. drummondii (Gulf of Mexico and Baja California).
In this study, we addressed the following questions: 1, Does the genetic diversity vary between populations? Because different environmental conditions allow different evolutionary pressures to act in each population, we expect variation in genetic diversity and related genetic parameters such as heterozygosity and inbreeding. 2, Does the predicted variation in genetic diversity relate to intrinsic biological properties of the populations, such as selfing or changes in floral traits? If self-fertilization is the predominant mating system in the population, we expect populations to show smaller flower size, higher inbreeding, lower heterozygosity, and reduced genetic diversity (Ingvarsson, 2002). 3, Do populations closer to the center of the distribution have greater genetic diversity and less genetic differentiation than peripheral populations? Large distances between populations of species with large ranges would be barriers to gene flow (Lawrence & Fraser, 2020). Also, species living in coastal dunes commonly experience rapid cycles of colonization and extinction, and have associated bottleneck events or founder effects (Eckert et al., 2008). This effect would be exacerbated in peripheral populations (Lawrence & Fraser, 2020). We expect that populations at the edge of the distribution would have lower genetic diversity and higher differentiation than central populations.
Materials and methods
Study system
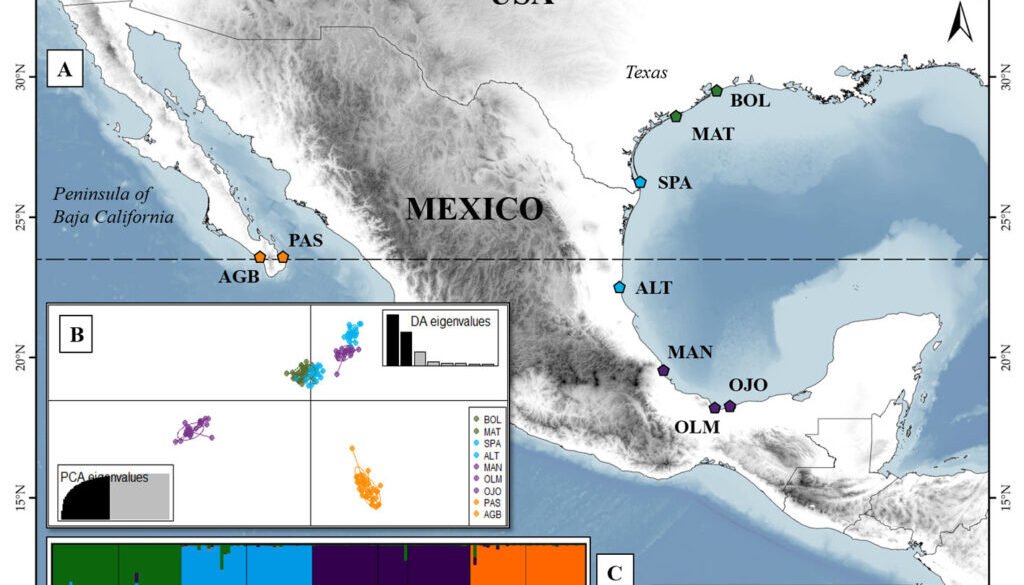
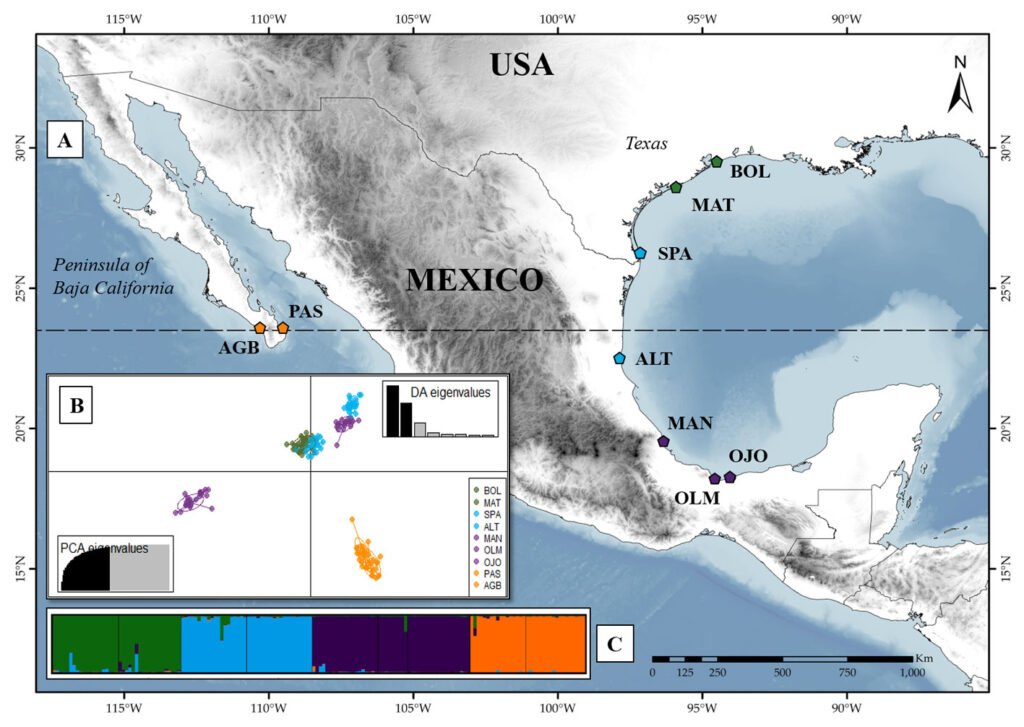
Oenothera drummondii is the sixth member of the series Raimannia within the subsection Raimannia in the genus Oenothera (Dietrich & Wagner, 1988). Its center of origin is in North America (Texas, USA) and consists of 2 subspecies with a disjunct distribution: O. drummondii subsp. drummondii, which is distributed in the coastal dunes of the Gulf of Mexico (from North Carolina in the USA to Campeche in Mexico), and O. drummondii subsp. thalasaphilla along the Pacific coast at the Southern tip of Baja California in Mexico (Dietrich & Wagner, 1988; Hernández-Espinosa et al., 2020). The 2 subspecies can be distinguished morphologically: O. drummondii subsp. drummondii have large flowers and long pubescent leaves, while O. drummondii subsp. thalasaphilla is characterized by having small flowers and small succulent leaves (Dietrich & Wagner, 1988). O. drummondii subsp. drummondii has been accidentally introduced into coastal dune systems around the world, and is considered invasive in several of them (Castillo-Infante et al., 2021; Dietrich & Wagner, 1988). The species is restricted to coastal dunes, inhabiting the dunes from the back beaches to the first dune ridges and embryo-dunes of the Gulf of Mexico and Southeastern USA (from Campeche to North Carolina) (Dietrich & Wagner, 1988; Gallego-Fernández & García-Franco, 2021b; Moreno-Casasola, 1988). From Louisiana to North Carolina in USA, O. drummondii shares its distribution with O. humifusa, another species within subsection Raimmania. Oenothera humifusa is an autogamous, permanent structural heterozygote (PTH), very similar to O. drummondii in growth form and habitat requirements. In the overlapped distribution, 2 basic types can be distinguished, although they are linked by intermediate forms that result from crosses between O. drummondii and O. humifusa (Dietrich & Wagner, 1988; Wagner et al., 2007). To ensure that the collection represented only O. drummondii and excluded the possibility of hybrids, the sampling was made systematically, beginning in Texas, USA, and continuing southward to Tabasco. Some of the populations collected were small and restricted to limited areas, in which all individuals were collected. The most illustrative example was the population of Ojoshal, composed of 9 individuals. We studied 169 individuals from 9 North American populations, and the 2 subspecies of O. drummondii (Fig. 1; Supplementary material: Table S1). Of these, 2 populations are from O. drummondii subsp. thalassaphila (Agua Blanca and Punta Arena del Sur), and the remaining 7 are from subsp. drummondii (Bolívar, South Padre Island, Matagorda, Altamira, La Mancha, Olmeca, and Ojoshal). We had information on self-compatibility and floral traits for 4 of the 9 populations evaluated (Bolívar, South Padre Island, La Mancha, and Ojoshal), since Gallego-Fernández and García-Franco (2021a, b) used the same populations in their studies (Supplementary material: Table S2).
Vegetal material
Leaf material was collected from reproductive adults of each population, for which 5 leaves per plant were sampled in paper bags with silica gel, and the samples were subsequently stored at -20 °C. When population density exceeded 30 individuals, 20 were randomly selected, and if within a population less than 20 individuals were found, as in Ojoshal, the entire population was sampled (Supplementary material: Table S1).
DNA extraction and microsatellite loci amplification
Ten microsatellite loci transferred to O. drummondii from other species of the Oenothera (Hernández-Espinosa et al., 2020) were used in this study: OenhaB105, OenhaD102, OenbidiA_C10, Oenbi2triA_A1, Oenbi2triA_D3, Oenbi39tri10, Oenbi2triA_H1, Oenbi39tri4, Oenbi2triA_E4 and Oenbi39di2. The extraction of genomic DNA was carried out according to 2 protocols: a) CTAB extraction, using 50 mg of dry leaf tissue (González & Vovides, 2002). The extracted DNA was subsequently purified using the PCR Clean-up & Gel Extraction Purification Kit (QIAGEN). b) DNeasy Plant Mini Kit using 20 mg dried leaf tissue following the manufacturer’s instructions (QIAGEN). The quantity and quality of the extracted DNA were verified on 1% agarose gels stained with Red Gel.
Microsatellite amplification was performed using 2 protocols. An initial PCR in a final volume of 10 µL containing total DNA, 2X reaction buffer, 0.2 mM of each dNTP, 1.6 mM MgCl2, 0.5 ng/µL BSA, 0.025 U/µL Taq polymerase, 0.5 µM reverse and forward primers, and 0.5 µM universal primer M13. For sample visualization, each forward primer was modified by the addition of an M13 sequence at the 5’ end (5’-TGT AAA ACG ACG GCC AGT-3’) that is complementary to an M13 primer labeled with either NED (yellow), HEX (green), or 6-FAM (blue) fluorophores. The amplification program consisted of an initial cycle of 3 min at 94 °C; followed by 25 cycles of 94 °C for 40 s, 50 °C for 40 s, and 72 °C for 60 s; 8 cycles of 94 °C for 60 s, 53 °C for 60 s, and 72 °C for 60 s, where hybridization of the M13 primer occurs; and a final extension at 72 °C for 10 min. A second PCR was performed using the Type-it microsatellite kit (QIAGEN), 1 µL of total DNA, and 0.2 µM of the forward, reverse, and M13 primers in cases where no amplification products were obtained from the first PCR after verification in 1% agarose. The amplification program consisted of a 15-minute cycle at 95 °C, followed by 20 cycles at 94 °C for 40 s, 50 °C for 90 s, and 72 °C for 60 s; 15 cycles at 94 °C for 60 s, 53 °C for 60 s, and 72 °C for 60 s. The final extension was performed at 60 °C for 30 min. A standard 400 bp marker and an ABI 3730 Gene Analysis System (Macrogen) were used to analyze the PCR products resulting from amplification. GeneMarker (Softgenetics, State College, PA, USA) was used to identify allele sizes manually. Oenothera drummondii is a diploid species, with amplifications consisting of 1 or 2 alleles per individual.
Statistical analysis
Presence of null alleles was evaluated using the FreeNA software with the EM algorithm (Chapuis & Estoup, 2007). The frequency of null alleles was estimated for each locus and population. FST values were calculated with FreeNA with the ENA algorithm for the correction of null alleles. The presence of clones in the populations was assessed with GenAIEx 6.503 software (Peakall & Smouse, 2012). Deviations from Hardy-Weinberg equilibrium for each locus in each population were assessed by the X2 test using the Benjamin-Hochberg procedure in GenAIEx 6.503 software (Peakall & Smouse, 2012). Linkage disequilibrium (LD) between loci was evaluated in Arlequin 3.5.2 (Excoffier & Lischer, 2010).
The percentage of polymorphic alleles (P), average number of alleles per locus (A), average number of effective alleles (Ae), observed heterozygosity (HO), expected heterozygosity (HE), and fixation index (F) were calculated in GenAIEx 6.503 (Peakall & Smouse, 2012). To estimate the patterns of genetic variation between and within the populations, we calculated the F-statistics (i.e., FIS, FIT and FST) (Wright, 1978). Additionally, FIS per population, considering null allele frequency, was estimated using INEST 2.2 (Chybicki, 2017), applying Bayesian method with 300,000 steps, sampling every 1,000 steps and burn-in of 30,000 steps. For the bottleneck analysis, the Wilcoxon signed rank test was performed for 3 mutation models: Infinite Allele Model (IAM), Stepwise Mutation Model (SMM), and Two-Phase Model (TPM). INEST 2.2 was used to run 100,000 simulations for each mutation. The relationship between latitude and parameters of genetic diversity (A, P, HO and HE) was evaluated using regression models (Sokal & Rohlf, 1995) in the R 4.2.0 program (R Core Team, 2020).
To estimate the molecular variation within and between populations, an analysis of molecular variance (AMOVA) was performed using 1,000 permutations in poppr package in R 4.2.0 software (R Core Team, 2020). To determine isolation by distance (IBD), a Mantel test (Mantel, 1967) was performed between the matrix of genetic differentiation (FST) with INA null allele correction, and population geographic distances (Euclidean) using 999 permutations in the vegan R package (R Core Team, 2020). We measured current dispersal rates (m) using BAYESASS, in which m is interpreted as the fraction of migrants per generation in one population that is derived from another population (Wilson & Rannala, 2003). The Bayesian clustering algorithm implemented in the Structure 2.3.4 software (Pritchard et al., 2010) was used to clarify the genetic structure of the samples by assigning individuals to genetic groups. The simulation was carried out with correlated allele frequencies and with mixture models according to ancestry. All analyses used 100,000 replicates, a Markov chain Monte Carlo (MCMC) burn-in period of 10,000 steps, and 10 replicates per K. The results obtained from Structure were processed with Structure Harvester (Earl & VonHoldt, 2012) to select the optimal K (highest value of ΔK) using the Evanno method (Evanno et al., 2005). Clumpak (Kopelman et al., 2015) and Distruct (Rosenberg, 2004) were used to visualize the bar graphs obtained in Structure. Discriminant analysis of principal components (DAPC) using the R package adegenet with 1,000 permutations for validation was also used to analyze the genetic structure of O. drummondii. It was based on the genetic distances (Cavalli-Sforza & Edwards, 1967) for each pair of populations, using the INA correction (Chapuis & Estoup, 2007) for the presence of null alleles.
Results
Frequency of null alleles
The 10 microsatellite loci amplified successfully in 161 individuals from 9 populations. Statistical tests of Hardy-Weinberg equilibrium for each locus in each population showed that most cases (50 of 78) did not deviate from equilibrium. For each population, the linkage disequilibrium analysis showed significant deviations in at least one locus. Mean null allele frequencies for all populations were low (0.06 ± 0.04). The mean null allele frequencies per population were moderate to low (0.01 ± 0.02-0.13 ± 0.14). Five loci exhibited high allele frequencies (> 0.2) in at least one population when analyzed across all population-locus combinations (Supplementary material: Tables S3, S4).
Genetic diversity
We detected 99 alleles for the 10 microsatellite loci. The total number of alleles detected had an average of 9.9 ± 5.6 alleles per locus, ranging from 4 for OenhaD102 to 23 for OenbidiA_C10. The level of polymorphism was high in all populations (mean 86.67 ± 14.14). The lowest percentage of polymorphism was found in Ojoshal (OJO), where 4 loci were monomorphic. The number of alleles per locus ranged from 1.80 in Ojoshal to 4.60 in Bolívar, with an overall mean number of alleles per locus of 2.28 ± 0.48. We found a total of 35 private alleles, ranging from 1 in La Mancha to 8 in Bolívar and Agua Blanca. The average effective number of alleles was 2.28 ± 0.48. It ranged from 1.58 in Olmeca to 3.06 in Bolívar (Table 1).
Expected heterozygosity ranged from 0.283 in Olmeca to 0.599 in Matagorda. The overall mean was 0.442 ± 0.104. The observed heterozygosity was also lowest in Olmeca (0.221), and highest in Matagorda (0.547). The mean total observed heterozygosity was 0.409 ± 0.097. The average observed heterozygosity was lower (HO < HE) than expected, indicating an overall homozygote excess. This is consistent with the results of the Hardy-Weinberg equilibrium test, which showed heterozygote deficiency for at least 1 locus in all populations. However, the Hardy-Weinberg equilibrium test also showed heterozygote excess in 4 populations: South Padre Island, Olmeca, Ojoshal, and Punta Arena del Sur. Of these, Ojoshal showed heterozygote excess at 5 of the 6 polymorphic loci (Table 1).
Table 1
Parameters of neutral genetic diversity of O. drummondii at 10 microsatellite loci. N = Sample size; P = percentage of polymorphic loci; A = mean allele number by locus; Ae = mean effective allele number; Ap = private alleles; HO and HE = mean observed heterozygosity, and mean expected heterozygosity; F = fixation index. Hardy-Weinberg equilibrium deviations (p < 0.05), test is the number of polymorphic loci evaluated, and (-) is the number of loci with deficiency and excess of heterozygotes (+).
| Population | N | P | A | Ae | Ap | Ho | HE | F | H-W deviations | ||
| Test | (+) | (-) | |||||||||
| Bolívar | 20 | 100 | 4.60 | 3.06 | 8 | 0.410 | 0.557 | 0.264 | 10 | – | 4 |
| Matagorda | 19 | 100 | 4.20 | 2.75 | 3 | 0.547 | 0.599 | 0.086 | 10 | – | 1 |
| South Padre Island | 20 | 80 | 3.60 | 2.54 | 2 | 0.385 | 0.433 | 0.110 | 8 | 1 | 1 |
| Altamira | 20 | 100 | 3.10 | 2.27 | 1 | 0.485 | 0.487 | 0.003 | 10 | – | 2 |
| La Mancha | 20 | 90 | 3.10 | 2.28 | 3 | 0.420 | 0.464 | 0.094 | 9 | – | 3 |
| Olmeca | 19 | 70 | 3.00 | 1.58 | 6 | 0.221 | 0.283 | 0.219 | 7 | 1 | 2 |
| Ojoshal | 9 | 60 | 1.80 | 1.61 | 0 | 0.489 | 0.301 | -0.623 | 6 | 5 | 1 |
| Agua Blanca | 17 | 90 | 4.10 | 2.31 | 8 | 0.324 | 0.435 | 0.256 | 9 | – | 4 |
| Punta Arena del Sur | 18 | 90 | 3.30 | 2.11 | 4 | 0.400 | 0.420 | 0.047 | 9 | 1 | 2 |
| Mean ± SD | 18 ± 3.54 | 86.7 ± 14.1 | 3.42 ± 0.83 | 2.28 ± 0.48 | 0.41 ± 0.097 | 0.44 ± 0.10 | 0.05 ± 0.27 |
Table 2
Bottleneck estimation for 9 populations of O. drummondii. Significant values are shown in bold. SSM: Step mutation model, TPM: two-phase model.
| Population | FIS (INest) | FIS (INest) 95% HDPI | Bottleneck test (p-value) | |
| SSM | TPM | |||
| Bolívar (BOL) | 0.2061 | 0.2061-0.3218 | 0.385 | 0.3473 |
| Matagorda (MAT) | 0.1221 | 0.0023-0.2274 | 0.0656 | 0.0421 |
| South Padre Island (SPA) | 0.0356 | 0.0011-0.0715 | 0.1913 | 0.1907 |
| Altamira (ALT) | 0.0051 | 0.0000-0.0049 | 0.0322 | 0.0245 |
| La Mancha (MAN) | 0.0214 | 0.0044-0.0568 | 0.0820 | 0.0486 |
| Olmeca (OLM) | 0.2446 | 0.2277-0.3479 | 0.9845 | 0.9845 |
| Ojoshal (OJO) | 0.0133 | 0.0000-0.0217 | 0.0264 | 0.0268 |
| Agua Blanca (AGB) | 0.0977 | 0.0586-0.1955 | 0.973 | 0.9625 |
| Punta Arena del Sur (PAS) | 0.0196 | 0.0000-0.471 | 0.5443 | 0.4556 |
The coefficients of inbreeding obtained using the INEST ranged from 0.0051 in Altamira to 0.2446 in Olmeca (Table 2). Three models were evaluated to detect the presence of recent genetic bottlenecks. The infinite allele model (IAM), the two-phase model (TPM), and the step mutation model (SSM). The SMM model has been suggested as a statistically conservative approach to detect microsatellite bottlenecks (Luikart et al., 1998). However, given the instability and wide range of mutation rates inherent to these markers, the TPM model may be the one that best explains the predominant mutation process at most loci (Di Rienzo et al., 1994). According to the TPM model, 4 populations in O. drummondii show possible recent bottlenecks: Altamira, Ojoshal, Matagorda and La Mancha (Table 2). Regression analyses revealed significant latitudinal clines in genetic diversity, with both alleles per locus (r = 0.69, p < 0.005) and expected heterozygosity (r = 0.65, p < 0.005) showing strong negative relationships with latitude. Latitude significantly predicted the percentage of polymorphic loci (r = 0.4, p < 0.05), whereas no significant relationship was observed for heterozygosity (Supplementary material: Table S5).
Genetic structure
When evaluating the distribution of genetic variation within and among populations, mean values of global inbreeding (FIT = 0.416 ± 0.163) and local inbreeding (FIS = 0.059 ± 0.195) indicated significant heterozygosity deficits. The mean value of genetic differentiation (FST) was 0.376 ± 0.132, indicating that 38% of the genetic variation in O. drummondii was due to differences between populations. Furthermore, genetic differentiation between pairs of populations was mostly high (> 0.15). Matagorda and Bolívar had the lowest differentiation (0.06), followed by Punta Arena del Sur and Agua Blanca (0.17), and Bolívar and South Padre Island (0.18). The analysis of molecular variance (AMOVA) showed that most genetic variation was found among individuals (54%), followed by that found among populations (38%) (Table 3).
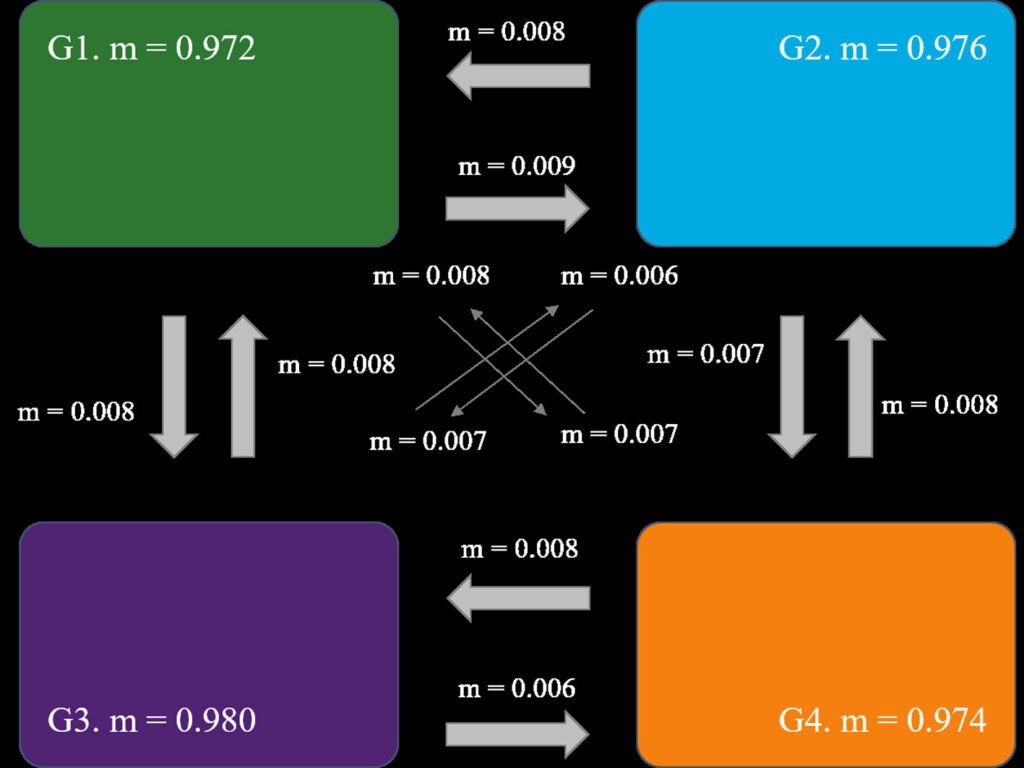
The Mantel test detected significant isolation by distance (r = 0.64, p = 0.002). The BAYESASS analysis revealed high diagonal m-values (G1 = 0.972 – G3 = 0.980), suggesting high gene flow within the individual groups. Higher gene flow was observed from G1 to G2 (m1 to 2 = 0.009) (Fig. 2). The results of the Bayesian clustering are shown in Supplementary material: Fig. S1. The most probable K with the highest value of ΔK = 479.023 was K = 3, followed by K = 4 with ΔK = 101.129. For K = 3, 3 clearly defined genetic groups were observed, with a very low proportion of admixture among them. In this case, the first group consisted of the Texas populations: Bolívar, Matagorda, and South Padre Island, and the northernmost Mexican population, Altamira. The second group included the 3 remaining Mexican populations in the Gulf of Mexico: La Mancha, Olmeca and Ojoshal, while the third group comprised the 2 populations on the Baja California Peninsula. For K = 4, the clustering is similar to K = 3. The main difference in this case was that the fourth cluster split the previous cluster containing the Texas and Altamira populations, indicating greater genetic correspondence between Bolívar and Matagorda (populations further north in the Gulf) and between South Padre Island and Altamira. Admixture was low, indicating that Oenothera populations are highly structured.
The Discriminant Analysis of Principal Components (DAPC), like the Bayesian method, also showed 4 groups (Fig. 1C; Supplementary material: Fig. S2). Consistent with the results of Structure, one group was formed by the populations of Baja California (Punta Arena del Sur and Agua Blanca). However, in this case, La Mancha was no longer grouped together with Ojoshal and Olmeca, which formed the second group. Altamira and La Mancha formed a third group that is closer to the fourth formed by the Texan populations (Bolívar, Matagorda, and South Padre Island). The DAPC results indicated the possibility that the Texas group was the ancestral group of the central Gulf of Mexico populations (Altamira and La Mancha). The other 2 groups were markedly more differentiated.
Table 3
Molecular variance analysis (AMOVA) of Oenothera drummondii populations.
| Source of variation | df | Sum sq | Mean sq | Variance component | Total variance (%) | p values |
| Between populations | 8 | 841.68 | 105.21 | 2.78 | 37.39 | 0.001 |
| Among populations | 153 | 807.84 | 5.28 | 0.61 | 8.21 | 0.001 |
| Within individuals | 162 | 657 | 4.05 | 4.05 | 54.39 | 0.001 |
| Total | 323 | 2,306.53 | 7.14 | 7.45 | 100 |
Discussion
In this research, we evaluated the genetic variation between and within populations of Oenothera drummondii throughout its range in North America, for which we used microsatellite markers. These markers demonstrated sufficient variability to resolve population genetic structure at high resolution, consistent with a previous study (Hernández-Espinosa et al., 2020). We also found that all populations were in linkage disequilibrium for at least one locus. Deviations may result from limited homologous recombination during meiosis when chromosomal rings are formed (Rauwolf et al., 2008). In O. drummondii, as in many Oenothera species, this limited recombination is reported to cause changes in the linkage equilibrium (Cleland, 1972; Rauwolf et al., 2008; Raven, 1979), as described in other ring-forming species of the genus: O. biennis (Larson et al., 2008; Levin, 1975; Levy & Levin, 1975), O. harringtonii (Skogen et al., 2012), O. hartwegii and O. gayleana (Lewis et al., 2016). The unique recombination system in the Oenothera is an attribute that has allowed them to colonize new environments from their origin in the Miocene (ca. 20 Ma), until recent times, successfully establishing themselves first in the Neotropics and then in Europe (Dietrich et al., 1997; Wagner et al., 2007).
Compared to other plants with similar characteristics in lifeform, reproductive system, geographic range, and dispersal mechanism, we found that Oenothera drummondii showed moderate to low genetic diversity. This is a short-lived perennial, self-compatible species with a mixed reproductive system and wide distribution (Dietrich & Wagner, 1988), and a variety of dispersal mechanisms (Gallego-Fernández et al., 2021). Mean levels of genetic diversity reported for other plants with mixed reproduction (N = 15, mean HE = 0.60), short-lived perennial life form (N = 29, mean HE = 0.55), wide distribution (N = 31, mean HE = 0.62), and water or wind dispersal (N = 28, mean HE = 0.61) (Nybom & Bartish, 2000), were higher than the level found in O. drummondii (mean HE = 0.442).
Historically, genetic diversity in Oenothera has been attributed to structural genomic rearrangements, particularly hybridization and reciprocal translocations that alter linkage relationships translocations (Cleland, 1972; Rauwolf et al., 2011), as the genomes in Oenothera were considered to be essentially “non-recombining” due to the formation of chromosomal rings during meiosis (Cleland, 1972; Golczyk et al., 2014; Rauwolf et al., 2008). In contrast, the reported genetic diversity in Oenothera species appears strongly influenced by their predominant reproductive system. O. biennis, a permanent translocation heterozygote (PTH), maintains higher genetic variation than O. drummondii, which exhibits a mixed mating system (mean HE = 0.69 vs. 0.44) (Larson et al., 2008). This agreed with the expectation of the “functional” asexual reproductive system of O. biennis, in which autogamy is almost complete and offspring are mostly clonal (Mather, 1943; Stebbins, 1957). Consistent with expectations for obligate sexual reproduction, self-incompatible, bivalent-forming Oenothera species (O. harringtonii with mean HE = 0.77, O. gayleana with mean HE = 0.53 and O. hartwegii with mean HE = 0.49 vs O. drummondii with HE = 0.44) maintained significantly higher genetic diversity than the mixed-mating O. drummondii (HE = 0.44) (Levin, 1975; Rhodes et al., 2014; Skogen et al., 2019).
In terms of its reproductive system, the self-compatible O. drummondii generally has large flowers and an elevated stigma, suggesting a high propensity for sexual reproduction, although it can self-pollinate in the absence of pollinators or in situations where self-fertilization is evolutionarily advantageous (Gregory, 1964; Sicard & Lenhard, 2011). Gallego-Fernández and García-Franco (2021b) assessed the level of self-compatibility of O. drummondii in the Gulf of Mexico, through germination essays. They evaluated 4 of the 9 populations in this study (Bolívar, South Padre Island, La Mancha and Ojoshal), and found that self-compatibility increased in low-latitude populations (La Mancha and Ojoshal), while lower for higher-latitude populations (Bolívar and South Padre Island). These results suggest a latitudinal shift towards greater selfing to the south of the distribution. It has been suggested that greater selfing in populations leads to a decrease in genetic diversity (Ingvarsson, 2002), which is consistent with the pattern of genetic variation found in this study. In agreement with the results of Gallego-Fernández and García-Franco (2021a), we found that populations with higher reported self-compatibility also had lower genetic diversity. On the other hand, in partial selfing species, reductions in genetic diversity are associated with changes in traits that facilitate selfing (i.e., reduced flower size, reduced nectar, pollen and reduced herkogamy) (Barrett & Harder, 1996; Shimizu & Tsuchimatsu, 2015; Tedder et al., 2015). According to the theory, it would be expected that the populations of O. drummondii with a higher self-compatibility, and lower genetic diversity (i.e., Ojoshal) would also show a tendency to reduce flower size, and populations with lower self-compatibility, and higher genetic diversity (i.e., Bolívar) would show an increase in flower size. However, Gallego-Fernández and García-Franco (2021a) found the opposite: populations showed larger flower sizes following a latitudinal gradient from subtropical (i.e., Bolívar and South Padre Island) to tropical (i.e., La Mancha and Ojoshal) climates. This result suggests that in O. drummondii there is no latitudinal correlation between a reduction in flower size and an increase in selfing, and thus to a reduction in genetic diversity. Differences in floral traits among O. drummondii populations appear to respond to both biotic (i.e., pollinators) and abiotic factors (i.e., temperature, precipitation), suggesting local adaptations to environmental changes (Gallego-Fernández & García-Franco, 2021a).
In O. drummondii the genetic diversity showed a significant increase with latitude, with its lowest values in Ojoshal and Olmeca, while the higher values were found in Bolívar and Matagorda, populations in the highest latitudes. The genus Oenothera appears to have originated in northern Mexico and the adjacent USA, and subsection Raimmania, to which O. drummondii belongs, is centered in this area (Dietrich & Wagner, 1988). Since older clades tend to have more genetic diversity (Willi et al., 2018), it would be expected that the greatest genetic diversity would be found in populations at latitudes closest to the center of origin (i.e., Bolívar and Matagorda) and that it would lower as populations moved away to the periphery of the distribution, as was found here. In addition to reducing population genetic variation through genetic drift, habitat suitability often decreases from the core to the edge of a species’ geographic range, leaving edge populations small and isolated (Sagarin & Gaines, 2002), as in Ojoshal. However, despite the small population size of Ojoshal, inbreeding was low, presumably because of its high selfing index (Gallego-Fernández & García-Franco, 2021b). Inbreeding depression is predicted to be lower in selfing species because recessive deleterious alleles are expected to be efficiently “purged” from populations (Winn et al., 2011), and selfing favors the selection of new recessive beneficial mutations (Burgarella & Glémin, 2017; Charlesworth et al., 1993).
The pattern of distribution of the genetic diversity found in Oenothera drummondii seems to be shaped by demographic and evolutionary processes (Durka, 1999; Hewitt, 1996). Historical climate-driven changes are known to still affect the present-day genetic diversity (Alvarez et al., 2009; Hewitt, 2000). As a result of sequential founder events during post-glacial recolonization, contemporary populations show reduced genetic diversity and increased genetic differentiation, especially at range margins (Eckert et al., 2008; Sagarin & Gaines, 2002). The above pattern was found in 512 North American herbs, showing that the shifting and fragmentation of species’ geographical ranges in the past 20,000 years has played an important role in shaping the genetic variation of contemporary populations (Benavides et al., 2019).
Regarding the populations in Baja California (i.e., Agua Blanca and Punta Arena del Sur), the results of both genetic diversity and genetic structure showed that they formed an independent genetic group, as expected given that the populations belong to a different subsp. of O. drummondii. However, based on the genetic differences detected and the evidence of differences in life history, vegetative and floral characteristics observed in the field at the time of collection, it is suggested that O. drummondii subsp. thalassaphila is a distinct species from O. drummondii subsp. drummondii (Table 4). A detailed taxonomic study and additional genetic data from new populations of O. drummondii subsp. thalassaphila will be needed to support this.
Table 4
Distinctive morphological attributes between O.drummondii subsp. drummondii and O. drummondii subsp. thalassaphila. Data taken from Benavides et al. (2019).
| O.drummondii subsp. drummondii | O. drummondii subsp. thalassaphila | |
| Habit | Annual herb | Perennial sudshrub |
| Stem pubescence | Strigillose to villous | Strigillose |
| Glandular puberulent hairs | Present | Absent |
| Cauline leaves | 1-8cm, oblanceolate to obovate, densely villous, margins entire to remotely sinuate dentate | 1-4.5cm, oblong lanceolate to oblanceolate, strigillose, marginis entire to coarsely dentate |
| Floral tube | 2.5-5cm, strigillose to densely villous | 2-3.5cm, strigillose |
| Sepals | 2-3cm, strigillose to villous | 1.3-2.5cm, strigillose, occasionally red doted |
| Petal length | 2.5-4.5cm | 2-3.5cm |
| Capsule | 2.5-5.5cm, strigillose to villous | 2-4cm, strigillose |
| Seeds | 1.1-1.7mm | 1.5-2mm, smooth |
We conclude that the genetic diversity and genetic structure across the range of O. drummondii in North America suggest a latitudinal shift towards increased selfing and a decline in genetic diversity in the low-latitude populations. However, there is no positive relation between reductions in floral traits and increased selfing in the populations. Differences in floral traits appear to respond to biotic and abiotic factors, suggesting local adaptations to environmental changes. In O. drummondii, genetic parameters may reflect historical processes like climate-driven changes (i.e., the last glaciation), and patterns of genetic distribution associated with them, such as the center-peripheral hypotheses. The genetic attributes evaluated here suggest that populations of O. drummondii subsp. thalassaphila may represent a distinct species, but further evaluation will be required to confirm this hypothesis.
Acknowledgments
This study was supported by the Consejo Nacional de Ciencia y Tecnología (Conacyt) by a scholarship (2019-000037-02NACF-29365), as part of a doctoral thesis directed by Jorge González-Astorga. This study was also supported by the Ministerio de Economía y Competitividad, Spain (MINECO Project CGL2015-65058-R co-funded by FEDER). We sincerely thank the assistance in fieldwork to collect plant materials: Rusty Feagin, José García-Franco, Alejandro Espinosa de los Monteros and Anwar Medina-Villareal. We also thank Janet Nolasco-Soto for the assistance in laboratory methods, and two anonymous reviewers for their comments and observations on the manuscript.
References
Alvarez, N., Thiel-Egenter, C., Tribsch, A., Holderegger, R., Manel, S., Schönswetter, P. et al. (2009). History or ecology? Substrate type as a major driver of patial genetic structure in Alpine plants. Ecology Letters, 12, 632–640. https://doi.org/10.1111/j.1461-0248.2009.01312.x
Barrett, S. C. H., & Harder, L. D. (1996). Ecology and evolution of plant mating. Trends in Ecology & Evolution, 11, 73–79. https://doi.org/10.1016/0169-5347(96)81046-9
Benavides, E., Kuethe, J. R., Ortiz-Alcaraz, A., & De La Luz, J. L. L. (2019). Oenothera resicum (Onagraceae), a new species and the first record of the family from the Revillagigedo Archipelago, Mexico. Phytotaxa, 416, 59–66. https://doi.org/10.11646/phytotaxa.416.1.7
Burgarella, C., & Glémin, S. (2017). Population genetics and genome evolution of selfing species. In J. Wiley & Sons (Ed.), Encyclopedia of Life Sciences (pp. 1–8). Hoboken, NJ: Wiley. https://doi.org/https://doi.org/10.1002/9780470015902.a0026804
Castillo-Infante, F. R., Mendoza-González, G., Rioja-Nieto, R., & Gallego-Fernández, J. B. (2021). Range Shifts in the Worldwide expansion of Oenothera drummondii subsp. drummondii, a plant species of coastal dunes. Diversity, 13, 603. https://doi.org/10.3390/d13110603
Cavalli-Sforza, L. L., & Edwards, A. W. F. (1967). Phylogenetic analysis. Models and estimation procedures. American Journal of Human Genetics, 19, 233.
Chapuis, M. P., & Estoup, A. (2007). Microsatellite null alleles and estimation of population differentiation. Molecular Biology and Evolution, 24, 621–631. https://doi.org/10.1093/molbev/msl191
Charlesworth, B., Morgan, M. T., & Charlesworth, D. (1993). The effect of deleterious mutations on neutral molecular variation. Genetics (United States), 134. https://doi.org/10.1093/genetics/134.4.1289
Charlesworth, D. (2006). Evolution of plant breeding systems. Current Biology, 16, R726–R735. https://doi.org/10.1016/j.cub.2006.07.068
Chung, M. Y., Merilä, J., Li, J., Mao, K., López-Pujol, J., Tsumura, Y. et al. (2023). Neutral and adaptive genetic diversity in plants: an overview. Frontiers in Ecology and Evolution, 11, 1116814. https://doi.org/10.3389/fevo.2023.1116814
Chybicki, I. J. (2017). INEST 2.2: the user manual. Poland: Department of Genetics, Kazimierz Wielki University.
Cleland, R. E. (1972). Oenothera, cytogenetics and evolution. London, NY: Academic Press.
Devaux, C., Lepers, C., & Porcher, E. (2014). Constraints imposed by pollinator behaviour on the ecology and evolution of plant mating systems. Journal of Evolutionary Biology, 27, 1413–1430. https://doi.org/10.1111/jeb.12380
Di Rienzo, A., Peterson, A. C., Garza, J. C., Valdes, A. M., Slatkin, M., & Freimer, N. B. (1994). Mutational processes of simple-sequence repeat loci in human populations. Proceedings of the National Academy of Sciences, 91, 3166–3170. https://doi.org/10.1073/pnas.91.8.3166
Dietrich, W., & Wagner, W. L. (1988). Systematics of Oenothera Section Oenothera Subsection Raimannia and Subsection Nutantigemma (Onagraceae). Systematic Botany Monographs, 24, 1–91. https://doi.org/10.2307/25027713
Dietrich, W., Wagner, W. L., & Raven, P. H. (1997). Systematics of Oenothera section Oenothera subsection Oenothera (Onagraceae). Systematic Botany Monographs, 50, 1–234. https://doi.org/10.2307/25027870
Duminil, J., Hardy, O. J., & Petit, R. J. (2009). Plant traits correlated with generation time directly affect inbreeding depression and mating system and indirectly genetic structure. BMC Evolutionary Biology, 9, 177. https://doi.org/10.1186/1471-2148-9-177
Durka, W. (1999). Genetic diversity in peripheral and subcentral populations ofCorrigiola litoralis L.(Illecebraceae). Heredity, 83, 476–484. https://doi.org/10.1038/sj.hdy.6886000
Earl, D. A., & VonHoldt, B. M. (2012). STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources, 4, 359–361. https://doi.org/10.1007/s12686-011-9548-7
Eckert, C. G., Samis, K. E., & Lougheed, S. C. (2008). Genetic variation across species’ geographical ranges: the central-marginal hypothesis and beyond. Molecular Ecology, 17, 1170–1188. https://doi.org/10.1111/j.1365-294X.2007.03659.x
Evanno, G., Regnaut, S., & Goudet, J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology, 14, 2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier, L., & Lischer, H. E. L. (2010). Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10, 564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Gallego-Fernández, J. B., & García-Franco, J. G. (2021a). Floral traits variation in Oenothera drummondii subsp. drummondii across a wide latitudinal range of native and non-native populations. Flora, 280, 151851. https://doi.org/https://doi.org/10.1016/j.flora.2021.151851
Gallego-Fernández, J. B., & García-Franco, J. G. (2021b). Self-compatibility and reproductive success of Oenothera drummondii subsp. drummondii: is it similar between native and non-native populations? Diversity, 13, 431. https://doi.org/10.3390/d13090431
Gallego-Fernández, J. B., Martínez, M. L., García-Franco, J. G., & Zunzunegui, M. (2021). Multiple seed dispersal modes of an invasive plant species on coastal dunes. Biological Invasions, 23, 111–127. https://doi.org/10.1007/s10530-020-02359-6
Golczyk, H., Massouh, A., & Greiner, S. (2014). Translocations of chromosome end-segments and facultative hetero-
chromatin promote meiotic ring formation in evening primroses. The Plant Cell, 26, 1280–1293. https://doi.org/10.1105/tpc.114.122655
González, D., & Vovides, A. (2002). Low intralineage divergence in Ceratozamia (Zamiaceae) detected with nuclear ribosomal DNA ITS and chloroplast DNA trnL-F non-coding region. Systematic Botany, 27, 654–661. https://doi.org/10.1043/0363-6445-27.4.654
Gregory, D. P. (1964). Hawmoth pollination in the Genus Oenothera. Journal of Systematic and Evolutionary Botany, 5, 385–420.
Greiner, S., & Köhl, K. (2014). Growing evening primroses (Oenothera). Frontiers in Plant Science, 5, 80880. https://doi.org/10.3389/fpls.2014.00038
Hamrick, J., & Godt, M. (1996). Effects of life history traits on genetic diversity in plant species. Philosophical Transactions of the Royal Society of London. Biological Sciences, 351, 1291–1298. https://doi.org/10.1098/rstb.1996.0112
He, Z. Z., Stotz, G. C., Liu, X., Liu, J. J., Wang, Y. G., Yang, J. et al. (2024). A global synthesis of the patterns of genetic diversity in endangered and invasive plants. Biological Conservation, 291, 110473. https://doi.org/https://doi.org/10.1016/j.biocon.2024.110473
Hernández-Espinosa, R., González-Astorga, J., de los Monteros, A. E., Cabrera-Toledo, D., & Gallego-Fernández, J. B. (2020). Transferability of microsatellite markers developed in Oenothera spp. To the invasive species Oenothera drummondii Hook. (Onagraceae). Diversity, 12, 1–13. https://doi.org/10.3390/d12100387
Hewitt, G. (2000). The genetic legacy of the Quaternary ice ages. Nature, 405, 907–913. https://doi.org/10.1038/35016000
Hewitt, G. M. (1996). Some genetic consequences of ice ages, and their role in divergence and speciation. Biological Journal of the Linnean Society, 58, 247–276. https://doi.org/10.1111/j.1095-8312.1996.tb01434.x
Holsinger, K. E. (1991). Mass-action models of plant mating systems: the evolutionary stability of mixed mating systems. The American Naturalist, 138, 606–622. https://doi.org/10.1086/285237
Ingvarsson, P. K. (2002). A metapopulation perspective on genetic diversity and differentiation in partially self-fertilizing plants. Evolution, 56, 2368–2373. https://doi.org/10.1111/j.0014-3820.2002.tb00162.x
Johnson, M. T. J., Agrawal, A. A., Maron, J. L., & Salminen, J. P. (2009). Heritability, covariation and natural selection on 24 traits of common evening primrose (Oenothera biennis) from a field experiment. Journal of Evolutio-
nary Biology, 22, 1296–1307. https://doi.org/10.1111/j.14209101.2009.01747.x
Johnson, M. T. J., Fitzjohn, R. G., Smith, S. D., Rausher, M. D., & Otto, S. P. (2011). Loss of sexual recombination and segregation is associated with increased diversification in evening primroses. Evolution, 65, 2873–2889. https://doi.org/10.1111/j.1558-5646.2011.01378.x
Koelling, V. A., Hamrick, J. L., & Mauricio, R. (2011). Genetic diversity and structure in two species of Leavenworthia with self-incompatible and self-compatible populations. Heredity, 106, 310–318. https://doi.org/10.1038/hdy.2010.59
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A., & Mayrose, I. (2015). Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Molecular Ecology Resources, 15, 1179–1191. https://doi.org/10.1111/1755-0998.12387
Larson, E. L., Bogdanowicz, S. M., Agrawal, A. A., Johnson, M. T. J., & Harrison, R. G. (2008). Isolation and characterization of polymorphic microsatellite loci in common evening primrose (Oenothera biennis). Molecular Ecology Notes, 8, 80–81. https://doi.org/10.1111/j.1471-8286.2007.01983.x
Lawrence, E. R., & Fraser, D. J. (2020). Latitudinal biodiversity gradients at three levels: linking species richness, population richness and genetic diversity. Global Ecology and Biogeography, 29, 770–788. https://doi.org/https://doi.org/10.1111/geb.13075
Levin, D. A. (1975). Genic heterozygosity and protein polymorphism among local populations of Oenothera biennis. Genetics, 79, 477–491. https://doi.org/10.1093/genetics/79.3.477
Levin, D. A. (2012). Mating system shifts on the trailing edge. Annals of Botany, 109, 613–620. https://doi.org/10.1093/aob/mcr159
Levy, M., & Levin, D. A. (1975). Genic heterozygosity and variation in permanent translocation heterozygotes of
the Oenothera biennis complex. Genetics, 79, 493–512. https://doi.org/10.1093/genetics/79.3.493
Lewis, E. M., Fant, J. B., Moore, M. J., Hastings, A. P., Larson, E. L., Agrawal, A. A. et al. (2016). Microsatellites for Oenothera gayleana and O. hartwegii subsp. filifolia (Onagraceae), and their Utility in section Calylophus. Applications in Plant Sciences, 4, 1500107. https://doi.org/10.3732/apps.1500107
Luikart, G., Allendorf, F., Cornuet, J., & Sherwin, W. (1998). Distortion of allele frequency distributions provides a test for recent population bolttlenecks. Heredity, 89, 238–247. https://doi.org/10.1093/jhered/89.3.238
Mantel, N. (1967). The detection of disease clustering and a generalized regression approach. Cancer Research, 27, 209–220.
Moreno-Casasola, P. (1988). Patterns of plant species distribution on coastal dunes along the Gulf of Mexico. Journal of Biogeography, 15, 787–806. https://doi.org/10.2307/2845340
Nybom, H., & Bartish, I. V. (2000). Effects of life history traits and sampling strategies on genetic diversity estimates obtained with RAPD markers in plants. Perspectives in Plant Ecology, Evolution and Systematics, 3, 93–114. https://doi.org/10.1078/1433-8319-00006
Opedal, Ø. H. (2018). Herkogamy, a principal functional trait of plant reproductive biology. International Journal of Plant Sciences, 179, 677–687. https://doi.org/10.1086/700314
Overson, R. P., Johnson, M. G., Bechen, L. L., Kinosian, S. P., Douglas, N. A., Fant, J. B. et al. (2023). A phylogeny of the evening primrose family (Onagraceae) using a target enrichment approach with 303 nuclear loci. BMC Ecology and Evolution, 23, 66. https://doi.org/10.1186/s12862-023-02151-9
Peakall, R., & Smouse, P. E. (2012). GenALEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics, 28, 2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Pelletier, T. A., & Carstens, B. C. (2018). Geographical range size and latitude predict population genetic structure in a global survey. Biology Letters, 14, 20170566. https://doi.org/10.1098/rsbl.2017.0566
Pritchard, J. K., Wen, X., & Falush, D. (2010). Documentation for structure software: Version 2.3. University of Chicago, Chicago.
R Core Team. (2020). R: a language and environment for statistical computing. R Foundation for Statistical Computing. https://www.r-project.org/
Raduski, A. R., Haney, E. B., & Igić, B. (2012). The expression of self-incompatibility in angiosperms is bimodal. Evolution, 66, 1275–1283. https://doi.org/10.1111/j.1558-5646.2011.01505.x
Rauwolf, U., Golczyk, H., Meurer, J., Herrmann, R. G., & Greiner, S. (2008). Molecular marker systems for Oeno-
thera genetics. Genetics, 180, 1289–1306. https://doi.org/10.1534/genetics.108.091249
Rauwolf, U., Greiner, S., Mráek, J., Rauwolf, M., Golczyk, H., Mohler, V. et al. (2011). Uncoupling of sexual reproduction from homologous recombination in homozygous Oenothera species. Heredity, 107, 87–94. https://doi.org/10.1038/hdy.2010.171
Raven, P. H. (1979). A survey of reproductive biology in Onagraceae. New Zealand Journal of Botany, 17, 575–593. https://doi.org/10.1080/0028825X.1979.10432572
Rhodes, M. K., Fant, J. B., & Skogen, K. A. (2014). Local topography shapes fine-scale spatial genetic structure in the Arkansas Valley evening primrose, Oenothera harringtonii (Onagraceae). Heredity, 105, 900–909. https://doi.org/10.5061/dryad
Rosenberg, N. A. (2004). DISTRUCT: a program for the graphical display of population structure. Molecular Ecology Notes, 4, 137–138. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Sagarin, R. D., & Gaines, S. D. (2002). The ‘abundant centre’ distribution: to what extent is it a biogeographical rule? Ecology Letters, 5, 137–147. https://doi.org/https://doi.org/10.1046/j.1461-0248.2002.00297.x
Shimizu, K. K., & Tsuchimatsu, T. (2015). Evolution of selfing: recurrent patterns in molecular adaptation.Annual Review of Ecology, Evolution, and Systematics, 46, 593–622. https://doi.org/10.1146/annurev-ecolsys-112414054249
Sicard, A., & Lenhard, M. (2011). The selfing syndrome: a model for studying the genetic and evolutionary basis of morphological adaptation in plants. Annals of Botany, 107, 1433–1443. https://doi.org/10.1146/annurev-ecolsys-112414-054249
Skogen, K. A., Hilpman, E. T., Todd, S. L., & Fant, J. B. (2012). Microsatellite primers in Oenothera harringtonii (Onagraceae), an annual endemic to the shortgrass prairie of Colorado. American Journal of Botany, 99, e313–e316. https://doi.org/10.3732/ajb.1200003
Skogen, K. A., Overson, R. P., Hilpman, E. T., & Fant, J. B. (2019). Hawkmoth pollination facilitates long-distance pollen dispersal and reduces isolation across a gradient of land-use change 1. Annals of the Missouri Botanical Garden, 104, 495–511. https://doi.org/10.3417/2019475
Sokal, R., & Rohlf, F. (1995). Biometry: the principles and practices of statistics in Biological Research (Third Ed.). New York: Freeman.
Stebbins, G. (1957). Self-fertilization and population variability in plants. The American Naturalist, 41, 337–352. https://doi.org/10.1086/281999
Tedder, A., Carleial, S., Gołębiewska, M., Kappel, C., Shimizu, K. K., & Stift, M. (2015). Evolution of the selfing syndrome in Arabis alpina (Brassicaceae). Plos One, 10, e0126618. https://doi.org/10.1371/journal.pone.0126618
Wagner, W. L., Hoch, P. C., & Raven, P. H. (2007). Revised classification of the Onagraceae. In C. Anderson (Ed.), Systematic Botany Monographs. The American Society of Plant Taxonomists. https://doi.org/10.1007/bf03027161
Whitehead, M. R., Lanfear, R., Mitchell, R. J., & Karron, J. D. (2018). Plant mating systems often vary widely among populations. Frontiers in Ecology and Evolution, 6, 38. https://doi.org/10.3389/fevo.2018.00038
Willi, Y., Fracassetti, M., Zoller, S., & Van Buskirk, J. (2018). Accumulation of Mutational Load at the Edges of a Species Range. Molecular Biology and Evolution, 35, 781–791. https://doi.org/10.1093/molbev/msy003
Wilson, G. A., & Rannala, B. (2003). Bayesian inference of recent migration rates using multilocus genotypes. Genetics, 163, 1177–1191. https://doi.org/10.1093/genetics/163.3.1177
Winn, A. A., Elle, E., Kalisz, S., Cheptou, P. O., Eckert, C. G., Goodwillie, C. et al. (2011). Analysis of inbreeding depression in mixed-mating plants provides evidence for selective interference and stable mixed mating. Evolution, 65, 3339–3359. https://doi.org/10.1111/j.15585646.2011.01462.x
Wright, S. (1969). Evolution and the genetics of populations: Vol. 2. The theory of gene frequencies. Chicago and
London: University of Chicago Press.
Wright, S. (1978). Evolution and the genetics of populations. Variability within and among natural populations. Vol. 4. Chicago, IL.: University of Chicago Press.