Primer genoma mitocondrial completo de un cangrejo tricodactílido: Rodriguezia adani (Brachyura: Trichodactyloidea: Trichodactylidae) de Mexico
Eric G. Moreno-Juáreza, b, *, Andrea Jiménez-Marínc, Fernando Álvareza
a Universidad Nacional Autónoma de México, Instituto de Biología, Colección Nacional de Crustáceos, Tercer Circuito s/n, Ciudad Universitaria, Coyoacán, 04510 Ciudad de México, Mexico
b Universidad Nacional Autónoma de México, Posgrado en Ciencias Biológicas, Instituto de Biología, Colección Nacional de Crustáceos, Tercer Circuito s/n, Ciudad Universitaria, Coyoacán, 04510 Ciudad de México, Mexico
c Universidad Nacional Autónoma de México, Instituto de Biología, Laboratorio de Biología Molecular, Tercer Circuito s/n, Ciudad Universitaria, Coyoacán, 04510 Ciudad de México, Mexico
*Corresponding author: ericgmorenoj@gmail.com (E.G. Moreno-Juárez)
Received: 13 August 2025; accepted: 01 December 2025
Abstract
We report for the first time the complete mitochondrial genome of a crab in the family Trichodactylidae, that of the stygobitic Rodriguezia adani Álvarez and Villalobos, 2018. The mitogenome is 17,561 bp of length with a GC composition of 26.4% and comprises 13 protein coding genes (PCGs), 2 ribosomal subunits and 19 transfer RNA (tRNAs) genes; its ribosomal subunits are separated by 10 genes (5 PCGs and 19 tRNAs). Its phylogenetic position was obtained through Maximum Likelihood and Bayesian Inference analyses, both placed this species inside the subsection Heterotremata and as the sister group of the family Orithyiidae. Changes in the number of tRNAs could be the result of the adaptation to a stygobitic lifestyle. This is the first genomic resource for the Trichodactylidae and could be the initial step for future phylogenomic studies of this group.
Keywords: Stygobitic; Freshwater crab; Cave; Mitochondrion
Resumen
Reportamos el primer genoma mitocondrial completo de un cangrejo de la familia Trichodactylidae, el de la especie estigobítica Rodriguezia adani Álvarez y Villalobos, 2018. El mitogenoma tiene 17,561 pb de longitud, una composición de GC de 26.4% y contiene 13 genes codificantes de proteínas (PCGs), 2 subunidades ribosomales y 19 genes de RNA de transferencia (tRNAs); sus subunidades ribosomales están separadas por 10 genes (5 PCGs y 19 tRNAs). La posición filogenética obtenida a través de máxima verosimilitud e inferencia bayesiana ubica a esta especie dentro de la subsección Heterotremata y como grupo hermano de la familia Orithyiidae. Los cambios en el número de tRNAs podrían ser el resultado de la adaptación a un estilo de vida estigobítico. Este es el primer recurso genómico para los Trichodactylidae y posiblemente el paso inicial para futuros estudios filogenómicos de este grupo.
Palabras clave: Estigobítico; Cangrejo dulceacuícola; Cueva; Mitocondria
Introduction
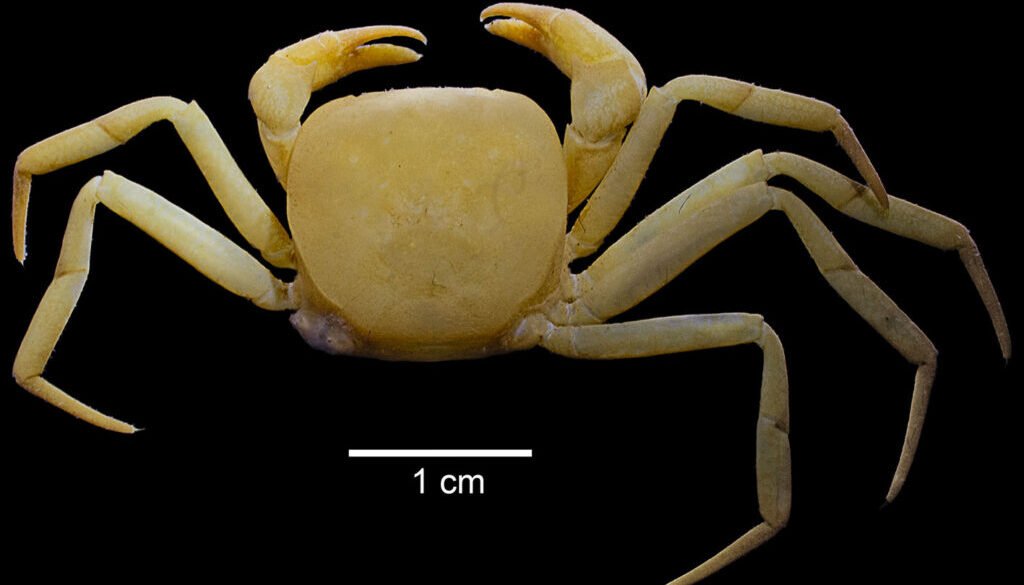
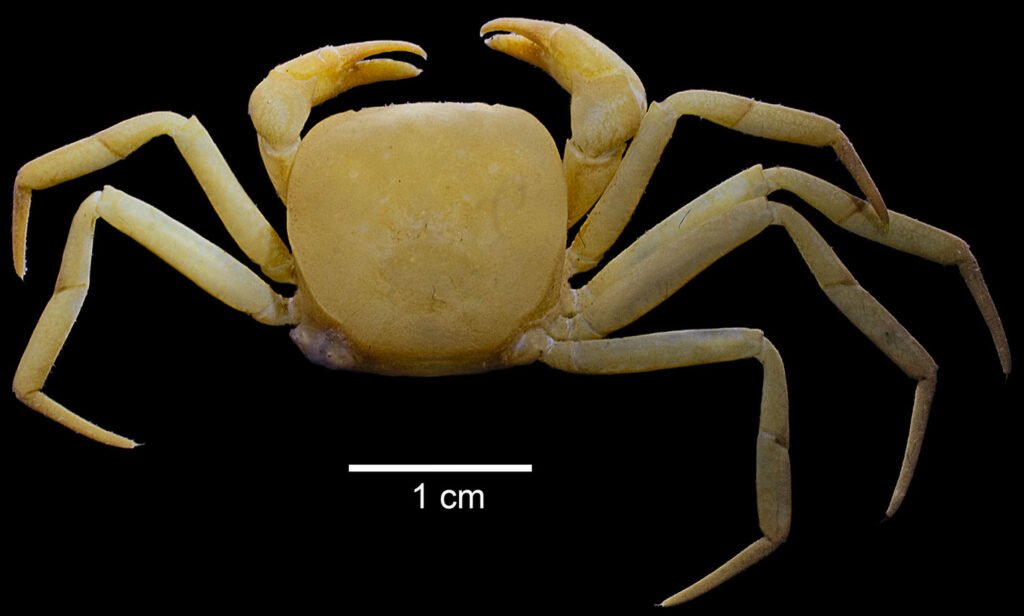
The family Trichodactylidae H. Milne-Edwards, 1853, the single family in the superfamily Trichodactyloidea, is one of the 3 families of freshwater crabs in the American Continent (Álvarez et al., 2020). Rodriguezia Bott (1969) one of the 2 genera of Trichodactylidae found in Mexico; these are primary freshwater crabs distributed in southern Mexico (Álvarez & Villalobos, 2018; Magalhães & Türkay, 1996). The genus Rodriguezia comprises 3 species, 2 of them stygobitic; from them, Rodriguezia adani Álvarez & Villalobos, 2018 was the last one to be described (Fig. 1). Most of the research on this family has focused on its distribution, taxonomy, ecology, conservation and anatomy (Collins et al., 2009, 2011; Cumberlidge et al., 2014; Lima-Gomes et al., 2017; Magalhães et al., 2016; Magalhães & Türkay, 1996; Weihrauch et al., 2004; Williner & Collins, 2013). However, few attempts have been made to understand its phylogenetic relationships, except for few molecular analyses performed to solve species complex issues in the genera Avotrichodactylus Pretzmann, 1968, Trichodactylus Latreille, 1828 and Dilocarcinus Milne Edwards, 1853 (Caetano-França et al., 2024; Ojeda et al., 2013; Souza-Carvalho et al., 2017). Studies using genomic sequencing for this family are lacking, only recently a study on the genome size of Trichodactylus fluviatilis Latreille, 1828 was published (Barioto et al., 2024). To further advance our knowledge on the phylogeny and evolution of the Trichodactylidae, we present the first complete mitochondrial genome for the family.

Materials and methods
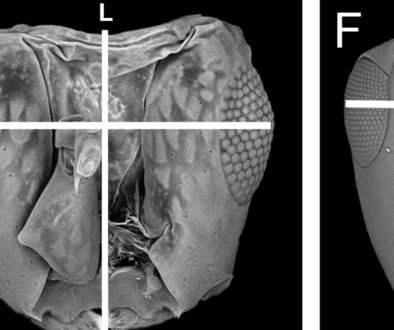
The organism used in this study was collected in the type locality, Aguas Blancas Cave, Tabasco, Mexico (17º36’22.06” N, 92º27’43.63” W, Fig. 2) and deposited in the Colección Nacional de Crustáceos (CNCR), of the Instituto de Biología, Universidad Nacional Autónoma de México (IB-UNAM), with catalog number CNCR 36762 (https://www.ib.unam.mx/ib/colecciones-biologicas/colecciones-zoologicas, Dr. Fernando Álvarez (falvarez@ib.unam.mx)). The organism was collected under license N° SGPA/DGVS/03775/22, provided by Semarnat. Its taxonomic identity was corroborated with Álvarez and Villalobos (2018) publication.
The DNA was purified from muscle tissue of the fifth pereopod with the Animal and Fungi DNA preparation kit, Jena Bioscience. Quality and concentration of DNA were determined with a NanoDrop 2000 spectrophotometer and Qubit 4 Fluorometer, respectively. The library used was prepared with Illumina DNA Prep Kit (Illumina, Inc.), following the Trevisan et al. (2019) protocol. The sequencing was conducted in an Illumina NextSeq 500 sequencer, using a 300-cycle cartridge in a Paired End (PE) 2X150 cycles configuration at the Massive Sequencing and Bioinformatic Unit of the Instituto de Biotecnología, UNAM.
A total of 11,090,166 raw sequences were obtained, its quality was measured with FastQC v. 0.11.9 (Andrews, 2010), and low quality reads (Q < 28) and adapters were removed with Trimmomatic v. 0.39 (Bolger et al., 2014). SPAdes v 3.15,1 (Bankevich et al., 2012) was implemented as assembler on Galaxy (The Galaxy Community, 2022) with k-mer values of 33, 55, 77, 105. The final sequence assembled was analyzed with MITOS (Bernt et al., 2013) to obtain the gene order and annotation. The framework of the Protein Coding Genes (PCGs) was corroborated with Geneious v. 2024.4 (Biomatters Inc., Auckland, New Zealand). To discard any issue during the assembly we performed the protocol of Vera-Paz et al. (2022). The depth average coverage was obtained with the online protocol of Ni et al. (2023). The final gene map was obtained with Chloroplot (Zheng et al., 2020).
To corroborate the phylogenetic position of R. adani, we initially based our selection of taxa on Tsang et al. (2014) and Wolfe et al. (2023), whose partial sequence-based phylogenies recovered Trichodactylidae together with Orithyiidae, Chasmocarcinidae, and Bellidae as a sister group to all Heterotremata. However, the only available mitogenome from that group was Orythia sinica (Orithyiidae), so to give a general idea of the phylogenetic position of R. adani,we selected other available mitogenomes from families and superfamilies representative of Heterotremata, species belonging to primary families of freshwater crabs, species of Thoracotremata, and Ranina ranina (Podotremata) as an outgroup (Table 1).
Table 1
Species included in the phylogenetic analysis and GenBank accession numbers.
| Species | Accession number | Reference |
| Rodriguezia adani | PQ064124 | This study |
| Ranina ranina | AB752308 | Unpublished |
| Tzotzilthelphusa villarosalensis | OP295767 | Moreno-Juárez et al., 2023 |
| Longpotamon yangtsekiens | KY785879 | Yuhui et al., 2017 |
| Ocypode ceratophthalmus | LN611669 | Tan et al., 2014 |
| Eriocheir japonica | FJ455505 | Wang et al., 2016 |
| Pachygrapsus crassipes | KC878511 | Yu et al., 2014 |
| Orithyia sinica | MG840649 | Zhong et al., 2018 |
| Chionoecetes japonicus | MT750295 | Kim et al., 2020 |
| Maguimithrax spinosissimus | KM405516 | Márquez et al., 2016 |
| Maja crispata | KY650651 | Basso et al., 2017 |
| Maja squinado | KY650652 | Basso et al., 2017 |
| Segonzacia mesatlantica | KY541839 | Mandon et al., 2017 |
| Austinograea alayseae | KC851803 | Kim et al., 2014 |
| Gandalfus puia | KR002727 | Kim et al., 2016 |
| Echinoecus nipponicus | MG574831 | Lee et al., 2018 |
| Pilumnus vespertilio | MF457402 | Tan et al., 2018 |
| Atergatis floridus | MG792341 | Karagozlu, Barbon et al., 2018 |
| Etisus anaglyptus | MG751773 | Karagozlu, Dihn et al., 2018 |
| Epixanthus frontalis | MF457404 | Tan et al., 2018 |
| Leptodius sanguineus | KT896744 | Sung et al., 2016 |
| Myomenippe fornasiniiq | LK391943 | Tan et al., 2016a |
| Myra affinis | MW192449 | Zhang et al., 2022 |
| Pyrhila pisum | KU343210 | Park et al., 2017 |
| Ashtoret lunaris | LK391941 | Tan et al., 2016b |
| Matuta victor | MT416712 | Huang et al., 2021 |
| Calappa bilineata | MN562587 | Lu et al., 2020 |
| Chaceon granulatus | AB769383 | Zhang et al., 2020 |
| Ovalipes punctatus | MH802052 | Unpublished |
| Scylla olivacea | FJ827760 | Unpublished |
| Charybdis bimaculata | MG787408 | Liu et al., 2018 |
| Callinectes sapidus | AY363392 | Place et al., 2005 |
| Portunus sanguinolentus | KT438509 | Ma et al., 2016 |
Table 2
Best partition scheme and model selection resulting from the analyses on IQ-Tree.
| Partition | Nucleotide substitution model | Reference |
| 12S | TVM + F + G4 | |
| 16S | TVM + F + I + G4 | |
| ATP6 pos1, COX2 pos1, COX3 pos1, CYTB pos1, ND3 pos1 | GTR + F + I + G4 | Tavaré, 1986 |
| ATP6 pos2, CYTB pos2, ND3 pos2 | GTR+F+I+G4 | Tavaré, 1986 |
| ATP6 pos3, ATP8 pos3, COX1 pos3, COX2 pos3, COX3 pos3, CYTB pos3, ND3 pos3 | GTR+F+I+G4 | Tavaré, 1986 |
| ATP8 pos1, ND2 pos1, ND6 pos1 | GTR+F+I+G4 | Tavaré, 1986 |
| ATP8 pos2, ND2 pos2, ND6 pos2 | GTR+F+I+G4 | Tavaré, 1986 |
| COX1 pos1 | TIM2+F+I+G4 | |
| COX1 pos2, COX2 pos2, COX3 pos2 | TVM+F+I+G4 | |
| ND1pos1, ND4 pos1, ND4L pos1, ND5 pos1 | GTR+F+I+G4 | Tavaré, 1986 |
| ND1 pos2, ND4 pos2, ND4L pos2, ND5 pos2 | GTR+F+I+G4 | Tavaré, 1986 |
| ND1 pos3, ND4 pos3, ND4L pos3, ND5 pos3 | TN+F+I+G4 | Tamura and Nei, 1993 |
| ND2 pos3, ND6 pos | TN+F+G4 | Tamura and Nei, 1993 |
A total of 15 mitochondrial genes (13 PCGs and 2 rRNA) were individually aligned with Mafft v. 7 online service (Katoh et al., 2019). The final concatenated matrix was analyzed with a codon partition merging and a model selection analysis, implemented on IQ-tree (Chernomor et al., 2016; Kalyaanamoorthy et al., 2017), resulting in 13 partitions and nucleotide substitution models (Table 2). Two phylogenetic inference methods were implemented, the maximum likelihood (ML) analysis was performed in IQ-Tree with 10,000 ultrafast bootstrap analysis (Hoang et al., 2018; Trifinopoulos et al., 2016). The Bayesian Inference (BI) analysis was performed with Exabayes (Aberer et al., 2014) with 20 million generations, sampling every 10,000 generations, standard deviation of split frequencies convergence value of 0.01 and 25% of burn-in.
Results
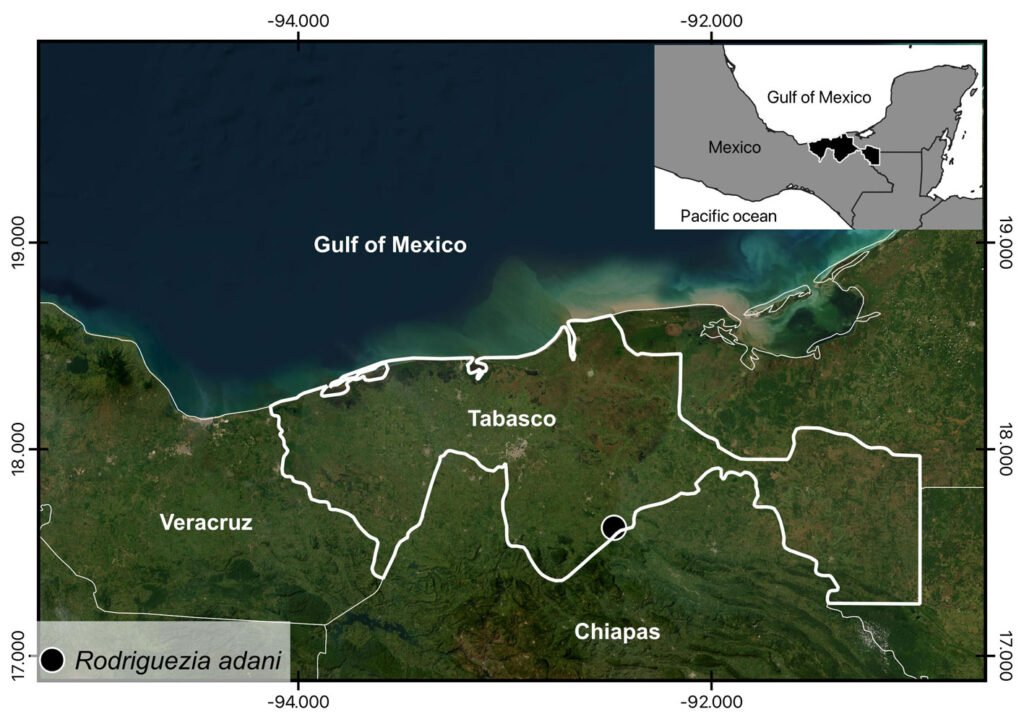
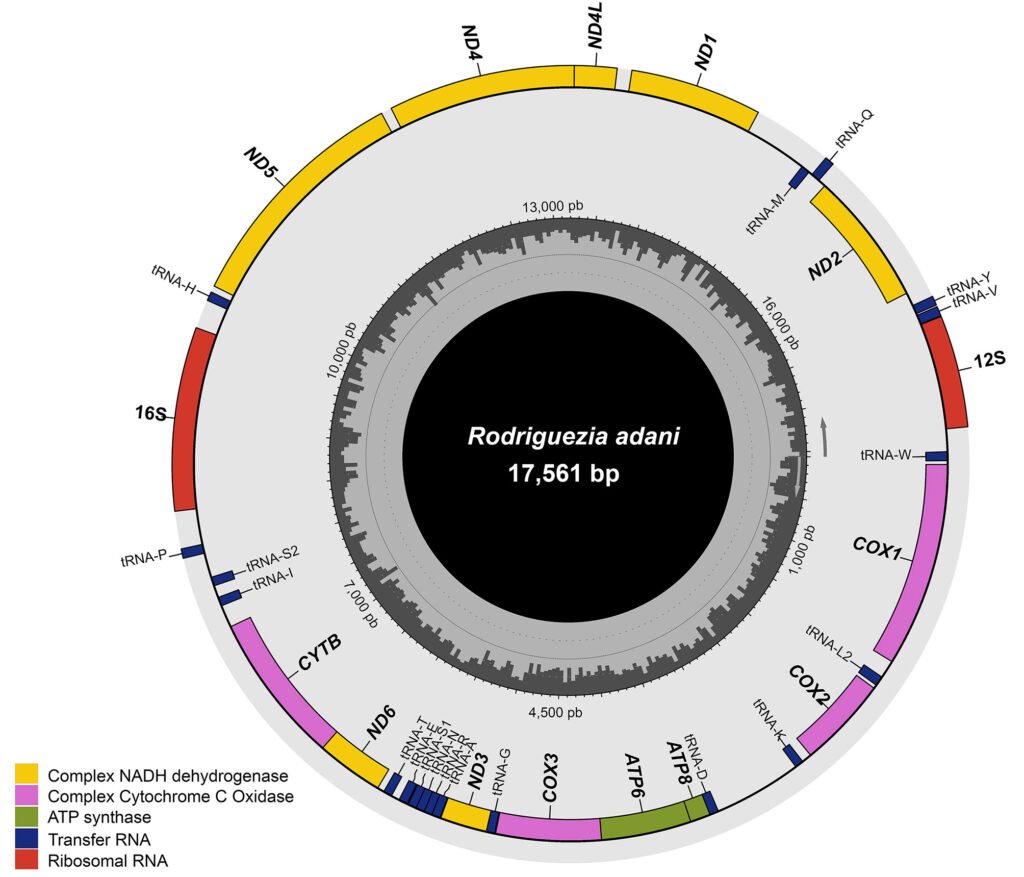
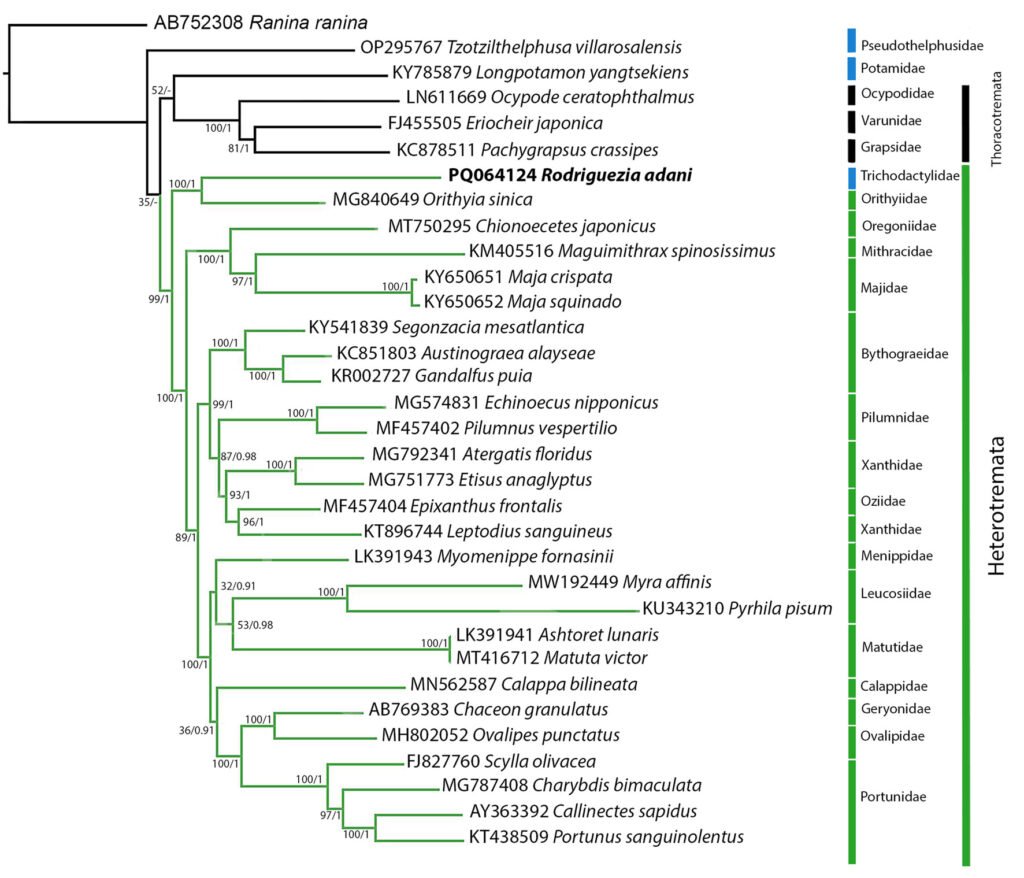
The complete mitochondrial genome of R. adani (GenBank: PQ064124; associated BioProject, SRA, and Bio-Sample numbers are PRJNA1139182, SRR29931217, and SAMN42764363, respectively) was obtained in only 1 contig with a length of 17,561 bp, an average depth of 125.8X and a GC composition of 26.4% (Fig. 3). It comprised 13 PCGs, 2 ribosomal RNA (rRNA) genes and 19 confirmed transfer RNA (tRNA) genes (Fig. 4, Table 3). Twenty-three genes are in the antisense strand: 9 PCGs (COX1, COX2, COX3, CYTB, ATP6, ATP8, ND1, ND3, ND6), and 14 tRNAs (tRNA-L1, K, D, G, A, R, N, S1, E, T, I, S2, M, W); and 11 are in the sense strand, 4 PCGs (ND1, ND4, ND4L, ND5), both rRNA genes (12S and 16S) and 5 tRNAs (tRNA-P, H, Q, Y, V); 1 tRNA-C is missing and tRNA-L2 and F are not confirmed. PCGs starting codons were ATG (COX1, COX2, COX3, ATP6, CYTB, ND2, ND4, ND4L), ATT (ATP8, ND5, ND6), ATC (ND3) and ATA (ND1). Most of the PCGs terminate with TAA, except for ND4 (TAG). Both phylogenetic analyses (ML, BI) recovered R. adani as the sister group of the family Orithyiidae with a high branch support (92/1) (Fig. 5).
Discussion
The family Trichodactylidae has been poorly studied molecularly and genomic sequencing of its species is non-existent, so this study contributes with the first mitochondrial genome sequenced from a stygobitic species. The gene order and topology found in R. adani is quite different from other crab mitochondrial genomes reported, this is due to the ribosomal subunits that are separated by several PCGs (5) and tRNAs (5). According to the mitochondrial patterns reported by Tan et al. (2019) this gene arrangement is not found in any other crab so far reported. Most decapod mitochondrial genomes present ribosomal subunits one next to the other or separated by tRNA-V or/and tRNA-Q, few decapod species (e.g., the crayfish Parastacus brasiliensis and Engaeus quadrimanus, reported in Tan et al., 2019) present ribosomal subunits separated by PCGs like R. adani. With respect to the tRNAs, although in mitochondrial genomes the most common number is 22, R. adani just presents 19 confirmed; several species could present more or less tRNAs, depending on their evolutionary history (Romanova et al., 2016, 2020, 2021; Tan et al., 2019). In the case of R. adani, its stygobitic condition could be related to physiological adaptations that enable survival in low food availability conditions and low oxygen concentrations (Hervant et al., 1999, 2001). It has been documented that in some instances metabolic changes can lead to the duplication or deletion of certain tRNAs, which can optimize the synthesis of enzymes or proteins that play an important role in these new metabolic processes (Romanova et al., 2021; Tan et al., 2019; Xing et al., 2025). Regarding tRNAs-L2 and F, both were recognized by MITOS and tRNAscan-SE but not supported by BlastN. It is possible that these tRNAs sequences are so novel that there is no sequence to compare them with, or an error may have occurred during assembly or annotation; for these reasons we decided to leave them as unconfirmed until other mitogenomes of the family are reported to corroborate or invalidate their presence through Romanova et al. (2016, 2020) protocol.
Table 3
Comparative table of the differences between R. adani and other brachyuran species.
| Organism | Size | Gene number | tRNAs | GC % composition | Reference |
| Rodriguezia adani | 17,561 | 34 | 19 | 26.4 | This study |
| Orithya sinica | 15,568 | 37 | 22 | 30.5 | Zhong et al., 2018 |
| Chionoecetes japonicus | 16,060 | 37 | 22 | 28.2 | Kim et al., 2020 |
| Callinectes sapidus | 16,263 | 37 | 22 | 30.9 | Place et al., 2005 |
| Eriocheir japonica | 16,352 | 37 | 22 | 28.4 | Wang et al., 2016 |
| Maja crispata | 16,592 | 37 | 22 | 29.7 | Basso et al., 2017 |
| Myra affinis | 15,349 | 37 | 22 | 29.4 | Basso et al., 2017 |
| Gandalfus puia | 15,548 | 37 | 22 | 30.1 | Kim et al., 2016 |
| Pilumnus vespertilio | 16,222 | 37 | 22 | 28.9 | Tan et al., 2018 |
| Epixanthus frontalis | 15,993 | 37 | 22 | 34.1 | Tan et al., 2018 |
| Matuta victor | 15,782 | 37 | 22 | 29.9 | Huang et al., 2021 |
| Calappa bilineata | 15,606 | 37 | 22 | 31.3 | Lu et al., 2020 |
| Ovalipes punctatus | 16,084 | 37 | 22 | 31.9 | Unpublished |
Regarding the phylogenetic position of R. adani, it was recovered as the sister group of the Orithyiidae, forming a clade that is the sister group to the rest of the subsection Heterotremata (Fig. 4).
Our results coincide with those of Wolfe et al. (2023), who using sequences of 10 genes, produced a phylogeny where the Orithyiidae and the Trichodactylidae are described as sister groups. Other coincidences with Wolfe et al. (2023) phylogenetic tree are the polyphyletic origin of primary freshwater crabs, Potamidae related to Thoracotremata, while Oregoniidae, Mithracidae, and Majidae are grouped in Majoidea and Portunidae, Ovalipidae and Geryonidae are grouped in Portunoidea. The phylogenetic relationships of the remaining species were not the aim of this work.
The low branch supports for these clades, likely due to a limited taxon sampling, preclude further interpretation. Future studies with more comprehensive sampling as in Tsang et al. (2014), Tan et al. (2019) or Wolfe et al. (2023), are needed to resolve these relationships with confidence (Nabhan & Sakar, 2012; Som, 2015).
Acknowledgements
We thank Ricardo Grande and Alejandro Sánchez for their assistance and advise during the genomic libraries’ construction and sequencing at the Unidad Universitaria de Secuenciación Masiva y Bioinformática, Instituto de Biotecnología, UNAM. This paper is part of the requirements for the Doctoral degree at Posgrado en Ciencias Biológicas, UNAM for EGMJ. EGMJ thanks Consejo Nacional de Humanidades, Ciencias y Tecnologías for the doctoral scholarship granted 2020-000013-01NACF-12928. Óscar Pérez Flores offered important advice during the phylogenetic analyses, which are greatly appreciated. Sandra Vera offered advice during the assembly and annotation which are greatly appreciated. José Luis Villalobos Hiriart, Kevin Madrigal and Josselyne Santillán for their help in the collection of the specimen; to the reviewers Ricardo Grande, Marco Solano and Carlos Pedraza for their interest in this work and their valuable comments. Finally, we thank Cristian Cervantes from the UniSSE, IB-UNAM. Funds were provided by Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica, Dirección General de Asuntos del Personal Académico, Universidad Nacional Autónoma de México grant IV200122 awarded to FA.
References
Aberer, A. J., Kobert, K., & Stamatakis, A. (2014). ExaBayes: massively parallel bayesian tree inference for the whole-genome era. Molecular Biology and Evolution, 31, 2553–2556. https://doi.org/10.1093/molbev/msu236
Álvarez, F., Ojeda, J. C., Souza-Carvalho, E., Villalobos, J. L., Magalhães, C., Wehrtmann, I. S. et al. (2020). Revision of the higher taxonomy of Neotropical freshwater crabs of the family Pseudothelphusidae Ortmann, 1893, based on multi-gene and morphological analyses. Zoological Journal of the Linnean Society, 193, 973–1001. https://doi.org/10.1093/zoolinnean/zlaa162
Álvarez, F., & Villalobos, J. L. (2018). A new species of stygobitic freshwater crab of the genus Rodriguezia Bott, 1969 (Crustacea: Decapoda: Trichodactylidae) from Tabasco, Mexico. Zootaxa, 4378, 137–143. https://doi.org/10.11646/zootaxa.4378.1.10
Andrews, S. (2010). FastQC a quality control tool for high throughput sequence data. Babraham Bioinformatics
http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A. A., Dvorkin, M., Kulikov, A. S. et al. (2012). SPAdes. A new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology, 19, 445–477. https://doi.org/10.1089/cmb.2012.0021
Barioto, J. G., Moraes, I. R. R., & Castilho, A. L. (2024). A first report of the genome size for the freshwater-crab family Trichodactylidae (Decapoda: Brachyura). Journal of Crustacean Biology, 44, ruae035. https://doi.org/10.1093/jcbiol/ruae035
Basso, A., Babbucci, M., Pauletto, M., Riginella, E., Pataenello, T., & Negrisolo, E. (2017). The highly rearranged mitochondrial genomes of the crabs Maja crispata and Maja squinado (Majidae) and gene order evolution in Brachyura. Scientific Reports, 7, 4096. https://doi.org/10.1038/s41598-017-04168-9
Bernt, M., Donath, A., Jühling, F., Externbrink, F., Florntz, C., Fritzsch, G. et al. (2013). MITOS: Improved de novo metazoan mitochondrial genome annotation. Molecular Phylogenetics and Evolution, 69, 313–319. https://doi.org/10.1016/j.ympev.2012.08.023
Bolger, M. A., Lohse, M., & Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics, 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Caetano-França, N. F., Magalhães, C., & Mantelatto, F. L. (2024). Integrative approach revealing a species complex in the Neotropical freshwater crab Dilocarcinus septemdentatus (Herbst, 1783) (Decapoda: Trichodactylidae) with a description of a new species. Arthropod Systematics & Phylogeny, 82, 385–405. https://doi.org/10.3897/asp.82.e115268
Chernomor, O., von Haeseler, A., & Minh, B. Q. (2016). Terrace aware data structure for phylogenomic inference from supermatrices. Systematic Biology, 65, 997–1008. https://doi.org/10.1093/sysbio/syw037
Collins, P. A., Giri, F., & Williner, V. (2009). Range extension for three species of South American freshwater crabs (Crustacea: Decapoda: Trichodactylidae). Zootaxa, 1977, 49–54. https://doi.org/10.11646/zootaxa.1977.1.4
Collins, P. A., Giri, F., & Williner, V. (2011). Biogeography of the freshwater decapods in the La Plata Basin, South America. Journal of Crustacean Biology, 31, 179–191. https://doi.org/10.1651/10-3306.1
Cumberlidge, N., Álvarez, F., & Villalobos, J. L. (2014). Results of the global conservation assessment of the
freshwater crabs (Brachyura, Pseudothelphusidae and Trichodactylidae): The Neotropical region, with an update on diversity. Zookeys, 457, 133–157. https://doi.org/10.3897/zookeys.457.6598
Hervant, F., Mathieu, J., & Barré, H. (1999). Comparative study on the metabolic responses of subterranean and surface-dwelling amphipods to long-term starvation and subsequent refeeding. Journal of Experimental Biology, 202, 3587–3595. https://jeb.biologists.org/content/202/24/3587
Hervant, F., Mathieu, J., & Durand, J. (2001). Behavioural, physiological and metabolic responses to long-term starvation and refeeding in a blind cave-dwelling (Proteus anguinus) and a surface-dwelling (Euproctus asper) salamander. Journal of Experimental Biology, 204, 269–281. https://jeb.biologists.org/content/204/2/269
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q., & Vinh, L. S. (2018). UFBoot2: Improving the ultrafast bootstrap approximation. Molecular Biology and Evolution, 35, 518–522. https://doi.org/10.1093/molbev/msx281
Huang, L., Zhong, S., Liu, Y., Huang, G., & Chen, X. (2021). The complete mitochondrial genome of Matuta victor (Decapoda: Matutidae) from Beibu Bay. Mitochondrial DNA B, 6, 241–242. https://doi.org/10.1080/23802359.2020.1861999
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A., & Jermiin, L. S. (2017). ModelFinder: Fast model selection for accurate phylogenetic estimates. Nature Methods, 14, 587–589. https://doi.org/10.1038/nmeth.4285
Karagozlu, M. Z., Barbon, M. M., Dihn, T. D., Demayo, C. G., & Kim, C. B. (2018). Complete mitochondrial genome of xanthid crab Atergatis floridus (Linnaeus, 1767). Mitochondrial DNA B, 3, 202–203. https://doi.org/10.1080/23802359.2018.1437832
Karagozlu, M. Z., Dihn, T. D., Nguyen, V. Q., & Kim, C. B. (2018). Analysis of complete mitochondrial genome of Etisus anaglyptus (Arthropoda, Decapoda, Xanthidae) with phylogenetic consideration. Mitochondrial DNA B, 24, 278–279. https://doi.org/10.1080/23802359.2018.1443038
Katoh, K., Rozewicki, J., & Yamada, K. D. (2019). MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20, 1160–1166. https://doi.org/10.1093/bib/bbx108
Kim, S. J., Kim, H. S., & Ju, S. J. (2014). Mitochondrial genome of the hydrothermal vent crab Austinograea alayseae (Crustacea: Bythograeidae): genetic differences between individuals from Tofua Arc and Manus Basin. Mitochondrial DNA, 25, 251–252. https://doi.org/10.3109/19401736.2013.800489
Kim, S. J., Kim, H. S., & Ju, S. J. (2016). Complete mitochondrial genome of the blind vent crab Gandalfus puia (Crustacea: Bythograeidae) from the Tonga Arc. Mitochondrial DNA A, 27, 2719–2720. https://doi.org/10.3109/19401736.2015.1046162
Kim, Y. H., Kim, K. R., Park, J. Y., Sung, M. S., Yun, B. H., Jeon, A. et al. (2020). The complete mitochondrial genome of Chionoecetes japonicus (Crustacea: Decapoda: Majoidea). Mitochondrial DNA B, 5, 3524–3526. https://doi.org/10.1080/23802359.2020.1827061
Lee, S. H., Oh, T. K., & Shin, M. H. (2018). The complete mitochondrial genome of Echinoecus nipponicus Miyake, 1939 (Crustacea: Decapoda: Pilumnidae), a symbiont of sea urchins. Mitochondrial DNA B, 5, 38–39. https://doi.org/10.1080/23802359.2017.1419093
Lima-Gomes, R., Farias-Lima, J., & Magalhães, C. (2017). Description of ten additional ossicles in the foregut of the freshwater crabs Sylviocarcinus pictus and Valdivia serrata (Decapoda: Trichodactylidae). Zoologia, 34, e1373. https://doi.org/10.3897/zoologia.34.e13731
Liu, X., Yuan, Y. M., He, Y. L., Liu, S. H., Ji, X., Qin, Y. T. et al. (2018). Characterization of the complete mitochondrial genome sequence of Charybdis bimaculata and phylogenetic analysis. Mitochondrial DNA B, 3, 1163–1164. https://doi.org/10.1080/23802359.2018.1524273
Lu, X., Gong, L., Zhang, Y., Chen, J., Liu, L., Jiang, L. et al. (2020). The complete mitochondrial genome of Calappa bilineata: The first representative from the family Calappidae and its phylogenetic position within Brachyura. Genomics, 112, 2516–2523. https://doi.org/10.1016/j.ygeno.2020.02.003
Ma, H. Y., Ma, C. Y., Zhu, J. J., Ren, G. J., Wang, W., Chen, W. et al. (2016). Characterization of the complete mitochondrial genome and phylogenetic relationships of the three-spot swimming crab (Portunus sanguinolentus). Genetics and Molecular Research, 15, gmr.15038580. https://dx.doi.org/10.4238/gmr.15038580
Magalhães, C., Campos, M. R., Collins, P. A., & Mantelatto, F. L. (2016). Diversity, distribution and conservation of freshwater crabs and shrimps in South America. In T. Kawai, & N. Cumberlidge (Eds.), A global overview of the conservation of freshwater decapods crustaceans (pp.303–338). Cham: Springer.
Magalhães, C., & Türkay, M. (1996). Taxonomy of the Neotropical freshwater crab family Trichodactylidae I. The generic system with description of some new genera (Crustacea: Decapoda: Brachyura). Senckenbergiana Biologica, 75, 63–95.
Mandon, P., Aznar-Cormano, L., Hourdez, S., & Samadi, S. (2017). Assembly of the mitochondrial genome of the hydrothermal vent crab Segonzacia mesatlantica and detection of potential nuclear pseudogenes. Mitochondrial DNA B, 2, 291–293. https://doi.org/10.1080/23802359.2017.1318674
Márquez, E. J., Hurtado-Alarcón, J. C., Isaza, J. P., Alzate, J. F., & Campos, N. H. (2016). Mitochondrial genome of the Caribbean king crab Damithrax spinosissimus (Lamarck, 1818) (Decapoda: Majidae). Mitochondrial DNA A, 27, 1724–1725. https://doi.org/10.3109/19401736.2014.961140
Moreno-Juárez, E. G., Jíménez-Marín, A., Vera-Paz, S. I., & Álvarez, F. (2023). The complete mitochondrial genome of the freshwater crab Tzotzilthelphusa villarosalensis Villalobos & Álvarez, 2013 (Brachyura: Decapoda: Pseudothelphusidae: Raddausinae) from Mexico. Journal of Crustacean Biology, 43, 1–9. https://doi.org/10.1093/jcbiol/ruad033
Nabhan, A. R., & Sakar, I. N. (2012). The impact of taxon sampling on phylogenetic inference: a review of two decades of controversy. Briefings in Bioinformatics, 13, 122–134. https://doi.org/10.1093/bib/bbr014
Ni, Y., Li, J. L., Zhang, C., & Liu, C. (2023). Generating sequencing depth and coverage map for organelle genomes. protocols.io. Retrieved November 20, 2024 from: https://doi:10.17504/protocols.io.4r3l27jkxg1y/v1
Ojeda, J. C., Álvarez, F., & Villalobos, J. L. (2013). Revisión de los cangrejos de agua dulce de la familia Trichodactylidae de México. In M. López-Mejía, & L. M. Mejía-Ortiz (Eds.), La carcinología en México (pp. 23–36). Chetumal: Universidad de Quintana Roo.
Park, Y. J., Park, C. E., Jung, B. K., Ibal, J. C., Jung, Y. G., Hong, S. J. et al. (2017). The first complete mitochondrial genome sequence of the leucosiid crab Pyrhila pisum (Arthropoda, Decapoda, Leucosiidae). Mitochondrial DNA B, 2, 885–886. https://doi.org/10.1080/23802359.2017.1407717
Place, A. R., Feng, X., Steven, C. R., Fourcade, H. M., & Boore, J. L. (2005). Genetic markers in blue crabs (Callinectes sapidus): II. Complete mitochondrial genome sequence and characterization of genetic variation. Journal of Experimental Marine Biology and Ecology, 319, 15–27. https://doi.org/10.1016/j.jembe.2004.03.024
Romanova, E. V., Aleoshin, V. V., Kamaltynov, R. M., Mikhailov, K. V., Logacheva M. D., Sirotinina, E. A. et al. (2016). Evolution of mitochondrial genomes in Baikalian amphipods. BMC Genomics, 17, 1016. https://doi.org/10.1186/s12864-016-3357-z
Romanova, E. V., Bukin, Y. S., Kamaltynov, R. M., Mikhailov, K. V., Logacheva M. D., Aleoshin, V. V. et al. (2020). Hidden cases of tRNA gene duplication and remolding in mitochondrial genomes of amphipods. Molecular Phylogenetics and Evolution, 144, 106710. https://doi.org/10.1016/j.ympev.2019.106710
Romanova, E. V., Bukin, Y. S., Kamaltynov, R. M., Mikhailov, K. V., Logacheva, M. D., Aleoshin, V. V. et al. (2021). The mitochondrial genome of a freshwater pelagic amphipod Macrohectopus branickii is among the longest in metazoan. Genes, 12, 2030. https://doi.org/10.3390/genes12122030
Som, A. 2015. Causes, consequences and solutions of phylogenetic incongruence. Briefings in Bioinformatics, 16, 536–548. https://doi.org/10.1093/bib/bbu015
Souza-Carvalho, E. A., Magalhães, C., & Mantelatto, F. L. (2017). Molecular phylogeny of the Trichodactylus fluviatilis Latreille, 1828 (Brachyura: Trichodactylidae) species complex. Journal of Crustacean Biology, 37, 187–194. https://doi.org/10.1093/jcbiol/rux005
Sung, J. M., Lee, J. H., Kim, S. G., Karagozlu, M. Z., & Kim, C. B. (2016). Complete mitochondrial genome of Leptodius sanguineus (Decapoda, Xanthidae). Mitochondrial DNA B, 1, 500–501. https://doi.org/10.1080/23802359.2016.1192505
Tamura, K., & Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution, 10, 512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Tan, M. H., Gan, H. M., Lee, Y. P., & Austin, C. M. (2014). The complete mitogenome of the ghost crab Ocypode ceratophthalmus (Pallas, 1772) (Crustacea: Decapoda: Ocypodidae). Mitochondrial DNA A, 27, 2123–2124. https://doi.org/10.3109/19401736.2014.982587
Tan, M. H., Gan, H. M., Lee, Y. P., & Austin, C. M. (2016a). The complete mitogenome of the stone crab Myomenippe fornasinii (Bianconi, 1851) (Crustacea: Decapoda: Menippidae). Mitochondrial DNA A, 27, 1374–1375. https://doi.org/10.3109/19401736.2014.947587
Tan, M. H., Gan, H. M., Lee, Y. P., & Austin, C. M. (2016b). The complete mitogenome of the moon crab Ashtoret lunaris (Forskal, 1775), (Crustacea; Decapoda; Matutidae). Mitochondrial DNA A, 27, 1313–1314. https://doi.org/10.3109/19401736.2014.945572
Tan, M. H., Gan, H. M., Lee, Y. P., Linton, S., Grandjean, F., Bartholomei-Santos, M. L. et al. (2018). ORDER within the chaos: insights into phylogenetic relationships within the Anomura (Crustacea: Decapoda) from mitochondrial sequences and gene order rearrangements. Molecular Phylogenetics and Evolution, 127, 320–331. https://doi.org/10.1016/j.ympev.2018.05.015
Tan, M. H., Gan, H. M., Lee, Y. P., Bracken-Grissom, H., Chan, T. Y., Miller, A. D. et al. (2019). Comparative mitogenomics of the Decapoda reveals evolutionary heterogeneity in architecture and composition. Scientific Reports, 9, 10756 https://doi.org/10.1038/s41598-019-47145-0
Tavaré, S. (1986). Some probabilistic and statistical problems in the analysis of DNA sequences. Lectures on Mathematics in the Life Sciences, 17, 57–86.
The Galaxy Community. (2022). The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2022 update. Nucleic Acids Research, 50, W345–W351. https://doi.org/10.1093/nar/gkac247
Trevisan, B., Alcantara, D. M. C., Machado, D. J., Marques, F. P. L., & Lahr, D. J. G. (2019). Genome skimming is a low-cost and robust strategy to assemble complete mitochondrial genomes from ethanol preserved specimens in biodiversity studies. PeerJ, 7, e7543. https://doi.org/10.7717/peerj.7543
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A., & Minh, B. Q. (2016). W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Research, 44, W232–W235. https://doi.org/10.1093/nar/gkw256
Tsang, L. M., Schubart, C. D., Ahyong, S. T., Lai, J. C. Y., Au, E. Y. C., Chan, T. Y. et al. (2014). Evolutionary history of true crabs (Crustacea: Decapoda: Brachyura) and the origin of freshwater crabs. Molecular Biology and Evolution, 31, 1173–1187. https://doi.org/10.1093/molbev/msu068
Vera-Paz, S. I., Contreras, D. D., Jost, M., Wanke, S., Rossado, A. J., Hernández-Guitíerrez, R. et al. (2022). New plastome structural rearrangements discovered in core Tillandsioideae (Bromeliaceae) support recently adopted taxonomy. Frontiers in Plant Science, 13, 924922. https://doi.org/10.3389/fpls.2022.924922
Wang, J., Huang, L., Cheng, Q., Lu, G., & Wang, C. (2016). Complete mitochondrial genomes of three mitten crabs, Eriocheir sinensis, E. hepuensis, and E. japonica. Mitochondrial DNA A, 27, 1175–1176. https://doi.org/10.3109/19401736.2014.936425
Weihrauch, D., Campbell, J., Towel, D. W., & Onken, H. (2004). Ion-motive ATPases and active, transbranchial NaCl uptake in the red freshwater crab, Dilocarcinus pagei (Decapoda,Trichodactylidae). Journal of Experimental Biology, 207, 4623–4631. https://doi.org/10.1242/jeb.01333
Williner, V., & Collins, P. A. (2013). Feeding ecology of the freshwater crab Trichodactylus borellianus (Decapoda: Trichodactylidae) in the floodplain of the Paraná River, southern South America. Latin American Journal of Aquatic Research, 41, 781–792. http://dx.doi.org/103856/vol41-issue4-fulltext-15
Wolfe, J., Ballou, L., Luque, J., Watson-Zink, V. M., Ahyong, S. T., Barido-Sottani, J. et al. (2023). Convergent adaptation of true crabs (Decapoda: Brachyura) to a gradient of terrestrial environments. Systematic Biology, 73, 247–262. https://doi.org/10.1093/sysbio/syad066
Xing, B., Lin, L., & Wu, Q. (2025). Application of mitochondrial genomes to species identification and evolution. Electronic Journal of Biotechnology, 75, 39–48. https://doi.org/10.1016/j.ejbt.2025.04.001
Yu, Y. Q., Ma, W. M., Yang, W. J., & Yang, J. S. (2014). The complete mitogenome of the lined shore crab Pachygrapsus crassipes Randall 1840 (Crustacea: Decapoda: Grapsidae). Mitochondrial DNA, 25, 263–264. https://doi.org/10.3109/19401736.2013.800497
Yuhui, X., Lijun, Z., Yue, H., Xiaoqi, W., Chen, Z., Huilun, Z. et al. (2017). Complete mitochondrial genomes from two species of Chinese freshwater crabs of the genus Sinopotamon recovered using next-generation sequencing reveal a novel gene order (Brachyura, Potamidae). Zookeys, 705, 41–60. https://doi.org/10.3897/zookeys.705.11852
Zhang, Y., Meng, L., Miao, Z., Wei, L., Liu, B., Liu, L. et al. (2022). Complete mitochondrial genome of Myra affinis (Decapoda: Brachyura: Leucosiidae) and its phylogenetic implications for brachyuran. Journal of Ocean University of China, 22, 987–997. https://doi.org/10.1007/s11802-022-4890-9
Zhang, Y., Wu, Y., Wang, X., Jiang, W., Yin, J., & Lin, Q. (2020). Comparative analysis of mitochondrial genome of a deep- sea crab Chaceon granulates reveals positive selection and novel genetic features. Journal of Oceanology and Limnology, 38, 427–437. https://doi.org/10.1007/s00343-019-8364-x
Zheng, S., Poczai, P., Hyvönen, J., Tang, J., & Amiryousefi, A. (2020). Chloroplot: an online program for the versatile plotting of organelle genomes. Frontiers in Genetics, 11, 576124. https://doi.org/10.3389/fgene.2020.576124
Zhong, S., Zhao, Y., & Zhang, Q. (2018). The first complete mitochondrial genome of Dorippoidea from Orithyia sinica (Decapoda: Orithyiidae). Mitochondrial DNA B, 3, 554–555. https://doi.org/10.1080/23802359.2018.1467237