Filogenia basada en COI y caracterización morfológica de una población brasileña de Lambornella trichoglossa (Ciliophora: Tetrahymenidae)
Vítor Ribeiro-Halfeld *
Universidade Federal de Juiz de Fora, Laboratório de Protozoologia, Rua José Lourenço Kelmer, s/n, São Pedro, Juiz de Fora, 36033-900 Minas Gerais, Brazil
*Corresponding author: ribeirohalfeldv@gmail.com (V. Ribeiro-Halfeld)
Received: 28 July 2025; accepted: 22 October 2025
Abstract
In this work, the morphological characterization and molecular phylogeny (SSU rDNA and COI) of Lambornella trichoglossa Foissner, 2003, found in phytotelmata environments in southeastern Brazil is presented. The morphological study performed with silver impregnation techniques demonstrated parameters similar to those recorded in the type population, and additional new information was also obtained, including the conjugation process and the occurrence of specimens with an increased number of caudal cilia. The phylogenetic reconstructions based on both markers suggest the inclusion of L. trichoglossa in the genus Tetrahymena Furgason, 1940. The COI sequence obtained in this work is the first mitochondrial sequence of the genus Lambornella deposited in GenBank and published in a scientific article.
Keywords: Tetrahymena; Phytotelmata; Molecular phylogeny
Resumen
En este trabajo se realizó la caracterización morfométrica y la filogenia molecular (SSU rDNA y COI) de Lambornella trichoglossa Foissner, 2003, encontrada en ambientes de fitotelmata en el sureste de Brasil. El estudio morfológico realizado con técnicas de impregnación de plata demostró parámetros similares a los registrados en la población tipo y también se obtuvo nueva información adicional, incluyendo el proceso de conjugación y la presencia de especímenes con un mayor número de cilios caudales. Las reconstrucciones filogenéticas basadas en ambos marcadores sugieren la inclusión de L. trichoglossa en el género Tetrahymena Furgason, 1940. La secuencia COI obtenida en este trabajo es la primera de tipo mitocondrial del género Lambornella depositada en GenBank y publicada en un artículo científico.
Palabras clave: Tetrahymena; Fitotelmas; Filogenia molecular
Introduction
The genus Lambornella has its systematic history marked by alterations. The type species, Lambornella stegomyiae Keilin, 1921, is a mosquito (Diptera, Culicidae) parasite. It was reclassified in the genus Tetrahymena Furgason, 1940 by Corliss (1960). However, the description of Lambornella clarki Corliss, 1976, another parasitic species, revalidated the genus, indicating the formation of cuticular cysts and the number of post-oral kineties as diagnostic features. Nevertheless, Strüder-Kypke et al. (2001) suggested the non-validity of the genus Lambornella once again, due to its position within the Tetrahymena clade, based on phylogenetic analyses inferred from small subunit ribosomal DNA (SSU). Results presented by Bourland and Stüder-Kypke (2010), and Dunthorn et al. (2012), also based on SSU sequences, demonstrated a similar grouping.
Lambornella trichoglossa Foissner, 2003 is the only free-living species of the genus. It occurs endemically in phytotelmata environments (Foissner, 2003). Despite the recurrent records in the Neotropical region (Buosi et al., 2014, 2015; Durán-Ramírez et al., 2015; Foissner, 2003; Foissner et al., 2003), no studies have investigated Lambornella’s phylogenetic position based on mitochondrial markers. According to Chantangsi and Lynn (2008), the mitochondrial gene cytochrome c oxidase-subunit I (COI) is effective in elucidating recent phylogenetic events in the genus Tetrahymena. In this context, the present work aimed to investigate the molecular phylogeny of L. trichoglossa based on the mitochondrial marker COI. Morphological analysis of the specimens was also performed in order to confirm specific identification and investigate possible morphological variation.
Materials and methods
One hundred milliliters of phytotelmata content were collected from 12 bromeliads belonging to Portea petropolitana at the Botanical Garden of the Federal University of Juiz de Fora (21°43’74” S, 43°22’06” W), in September, 2019. On the same day of collections, the samples were analyzed under a stereoscopic microscope with transmitted light. Active ciliates were picked with glass micropipettes and processed, according to Foissner (2014), to perform silver carbonate and dry silver nitrate impregnation techniques. From the collection day, over a period of 7 days, 20 specimens of L. trichoglossa were screened from the samples and measured in vivo to check for possible alterations in body length. Obtained data were analyzed using the Shapiro-Wilk normality test, and the means recorded on the first and seventh days were compared using the Student’s t-test. All statistical analyses related to the morphological data were conducted using PAST software, version 4.03 (Hammer et al., 2001).
Thirty specimens of L. trichoglossa were picked from the samples and fixed in absolute ethanol for molecular analysis. Total DNA extraction was performed using the Blood and Tissue kit (Qiagen®), following the manufacturer’s guidelines. Primers F388dT and R1184dT (Strüder-Kypke & Lynn, 2010) were used to amplify the COI gene in 25-microliter reactions. The SSU marker was also sequenced for comparison with the type population. Therefore, primers 18S F9Euk and 18S R1513 (Schrallhammer et al., 2013) were used.
PCR products were visualized in a 1% agarose gel and purified using the QIAquick PCR Purification Kit (Qiagen®), following the manufacturer’s guidelines. Subsequently, the material was sent for sequencing according to the Sanger method in 7 µl reactions, using the M13 forward and M13 reverse primers (Messing, 1983) for COI, and 18S R536, 18S F783, 18S F919, and 18S R1052 for SSU (Modeo et al., 2006). Sequencing reactions were performed on an ABI PRISM® 3100 sequencer (Applied Biosystems).
Obtained sequences were added to the respective datasets, jointly with COI and SSU sequences of approximately 20 species of the genus Tetrahymena, as well as outgroups obtained from GenBank accessed in May 2025. Sequences were aligned using MAFFT software, version 7 (Katoh et al., 2017). The resulting alignment was edited in the GBlocks platform, version 0.91b (Talavera & Catresana, 2007). The determination of the best nucleotide substitution model (GTR + G + I to COI, and TN93 + G to SSU) was performed with the aid of MegaX software (Kumar et al., 2018), using the maximum likelihood method, considering all sites. The maximum likelihood phylogenetic analysis was inferred using RAxML software, version 8 (Stamatakis, 2014), at its default settings, with 500 bootstrap replicates for COI, and 1,200 for SSU. The evolutionary distances of COI and SSU sequences were computed in Mega X, using the Kimura 2-parameter method (Kimura, 1980). A third dataset with 74 COI sequences of Tetrahymena species was prepared, in addition to outgroups, with the aim of performing the pairwise distance calculations with a greater number of species (Supplementary material). For this analysis, the Maximum Composite Likelihood model (Tamura et al., 2004) was used, with the aid of MegaX (Kumar et al., 2018).
Results
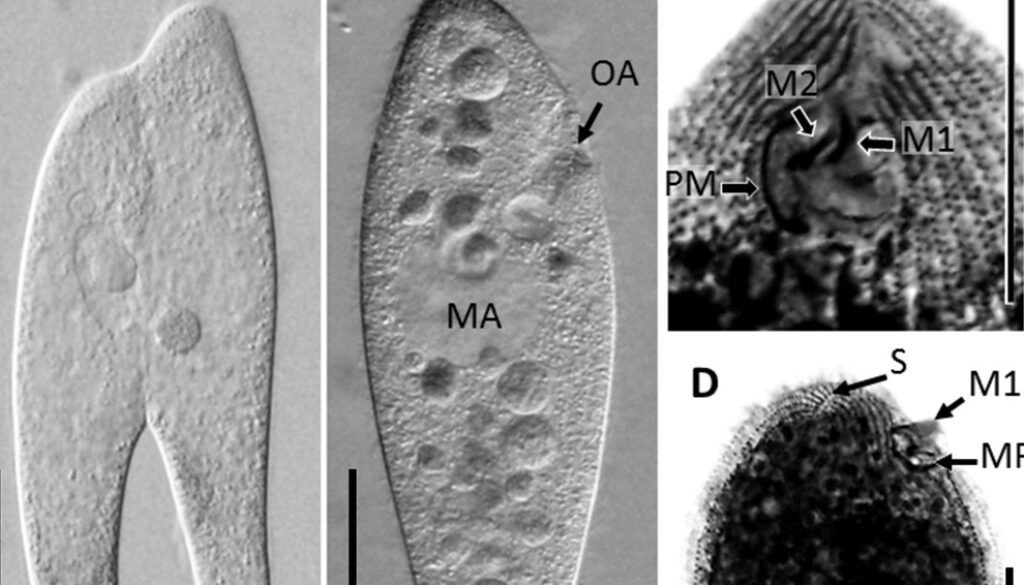
Table 1 presents the morphological data obtained. Morphological characterization of the L. trichoglossa population analyzed in this study resembles the type population described by Foissner (2003), presenting slightly higher values for body size and the number of somatic kineties. However, postoral kineties appear in lower numbers. Another important morphological characteristic distinct from the type population detected in the present study was the occurrence of specimens with a greater number of caudal cilia (Fig. 1h). These organisms have a complex of caudal cilia, as shown in Figure 1h and Supplementary material. However, they were not included in the morphological statistical analysis due to the unsatisfactory results of the impregnation techniques in their oral and some somatic structures. Opportunely, conjugating forms of L. trichoglossa were recorded, which is unprecedented for the species (Fig. 1a). The temporal analyses on body length performed demonstrated a significant decrease in the average size of the organisms over 7 days (Fig. 2, Table 2).
The obtained SSU sequence was deposited in GenBank with accession number MN567691. It matches 99.97% of the type population’s sequence (AJ810078) (Table 3). The phylogenetic reconstruction inferred by the maximum likelihood method revealed that the new sequence is grouped with L. trichoglossa (AJ810078), T. corlissi (U17356), and T. berger (AF364039) (Fig. 3).
Lambornella trichoglossa’s COI sequence was deposited in GenBank with accession number MN477017. It has 774 nucleotides, with the following percentage of bases: A = 33%, C = 12%, G = 13.4%, and T = 41.6%. Phylogenetic reconstruction using the maximum likelihood method demonstrated L. trichoglossa clustering in a clade with the same topology revealed by the SSU phylogeny; it also grouped with T. corlissi (EF070279) and T. bergeri (EF070270) with a support value of 86%, besides T. scolopax (KJ028669) (Fig. 4). The average paired difference of the 74 Tetraymena sequences used in the phylogenetic analyses was 12.99%. The L. trichoglossa sequence differs by 13.04% from the T. corlissi sequence (Table 4). Although the value is slightly above the average for the Tetrahymena species analyzed, this percentage is lower than the difference observed between T. corlissi and T. caudata (14.29%), T. paravorax (14%), and T. glochidiophila (16.39%), for example.
Table 1
Morphological data on silver carbonate-impregnated specimens of Lambornella trichoglossa.
| x | M | SD | SE | CV | Min | Max | n | |
| Body, lenght | 269.75 | 269.43 | 26.72 | 4.88 | 9.906 | 226.65 | 327.91 | 30 |
| Body, width in ventral view | 73.105 | 73.295 | 16.05 | 2.93 | 21.96 | 47.44 | 107.77 | 30 |
| Body lenght: Lateral width, ratio | 3.82 | 3.906 | 0.81 | 0.14 | 21.19 | 2.606 | 5.36 | 30 |
| Anterior body end to membranelle 1, distance | 47.91 | 48.73 | 7.407 | 1.35 | 15.46 | 34.14 | 63.87 | 30 |
| Oral opening, width | 32.45 | 31.42 | 5.11 | 0.93 | 15.77 | 21.28 | 44.12 | 30 |
| Anterior body end to macronucleus, distance | 94.55 | 92.88 | 13.86 | 2.53 | 14.66 | 74.16 | 126.35 | 30 |
| Macronucleus, length | 49.72 | 43.96 | 11.48 | 2.09 | 23.08 | 36.92 | 77.75 | 30 |
| Macronucleus, width | 51.27 | 48.71 | 9.42 | 1.72 | 18.38 | 33.74 | 66.86 | 30 |
| Macronucleus, number | 1 | 1 | 0 | 0 | 0 | 1 | 1 | 30 |
| Micronucleus, width | 5.67 | 5.5 | 1.15 | 0.21 | 20.408 | 4 | 9.56 | 27 |
| Micronucleus, number | 1.03 | 1 | 0.19 | 0.035 | 18.68 | 1 | 2 | 27 |
| Somatic kineties, total number | 52.33 | 52.5 | 3.79 | 0.69 | 7.24 | 45 | 62 | 30 |
| Somatic kineties, postoral number | 6.1 | 6 | 0.84 | 0.15 | 13.85 | 5 | 8 | 30 |
| Caudal cilia, number | 1 | 1 | 0 | 0 | 0 | 1 | 1 | 30 |
Measurements in μm. x, arithmetic mean. M, median. SD, standard deviation. SE, standard error of arithmetic mean. CV, coefficient of variation (%). Min, minimum. Max, maximum. n, number of individuals investigated.
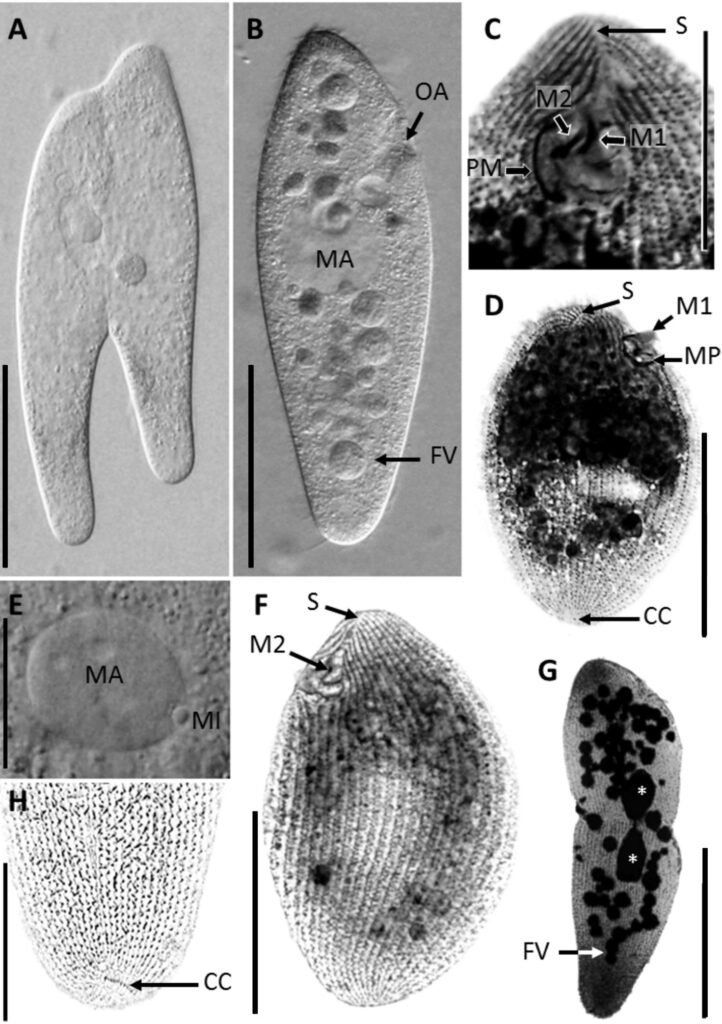
Figure 1. Morfological characterization of Lambornella trichoglossa. A, B, E. In vivo. C, D, F. Dry silver impregnation. G, H. Silver Carbonate impregnation. A, First recorded conjugating specimens; B, lateral view; C, oral apparatus; D, specimen with normal number of caudal cilium (dorsal view); E, nuclear apparatus; F, ventral view, showing oral apparatus and postoral kineties, below; G, specimen recorded during divisional morphogenesis, with multiple food vacuoles scattered throughout the body; H, detail of caudal cilia complex recorded in larger specimens. OA (oral apparatus), FV (food vacuoles), S (preoral suture), PM (paroral membrane), M1 and M2 (adoral membranelles), MA (macronucleus), MI (micronucleus), CC (caudal cilia). Asterisks indicate macronucleus during divisional morphogenesis process. Scale bars in A, B, D, F, G = 100 μm. In C, E H = 50μm.
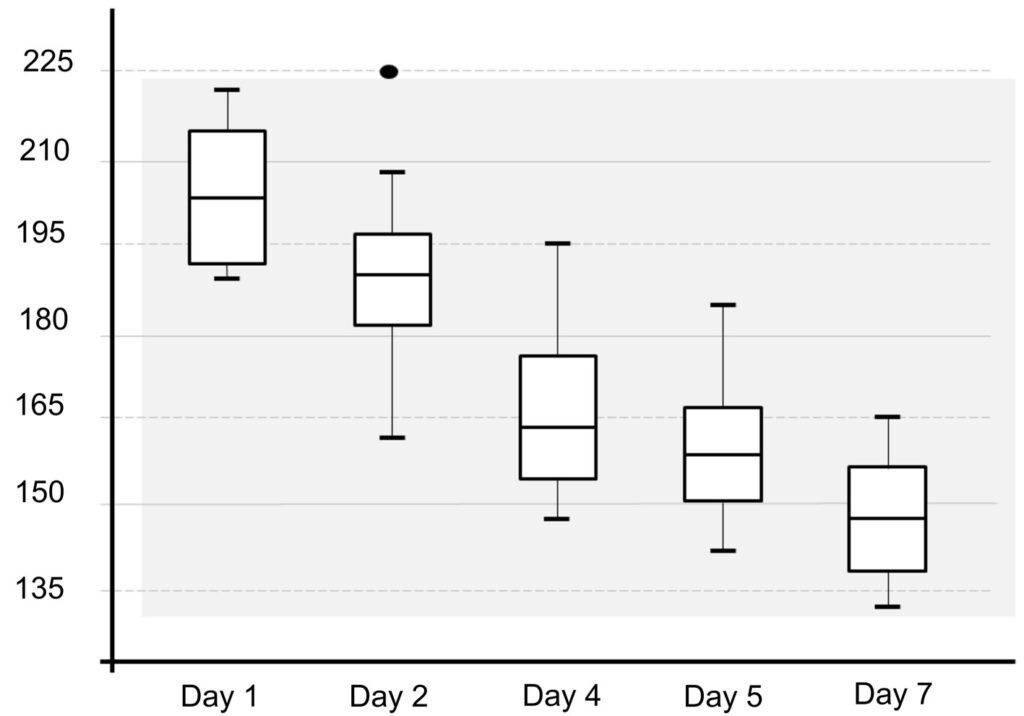
Figure 2. Body length reduction of L. trichoglossa over a 7-day period.
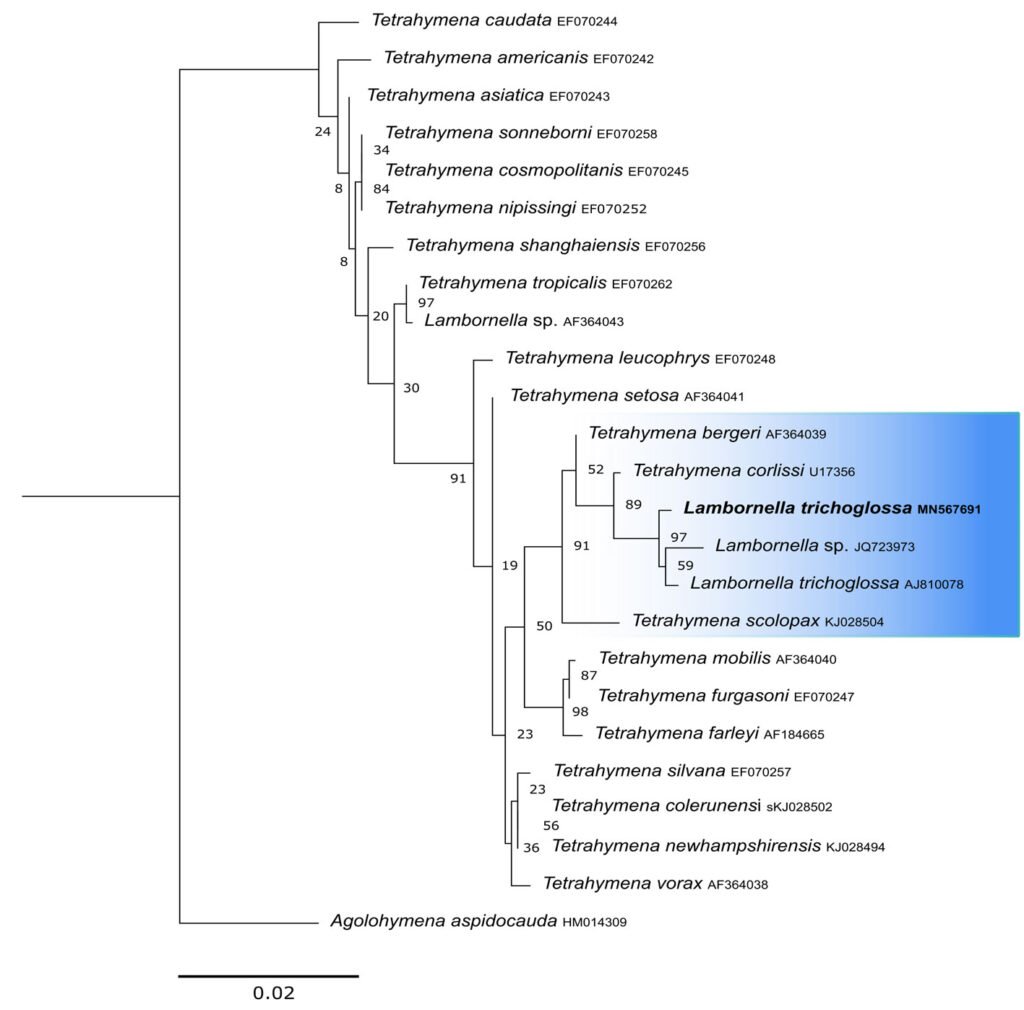
Figure 3. Maximum likelihood (ML) tree inferred from small subunit ribosomal DNA (SSU rDNA). Bold: sequence obtained in the present work. Numbers at nodes represent the bootstrap values of ML out of 1,200 replicates.
Table 2
Statistical data on the difference in body length of L. trichoglossa 7 days after collection.
| Number of measured specimens | |
| 1st day | 20 |
| 7th day | 20 |
| Body length mean (μm) | |
| 1st day | 205.66 |
| 7th day | 148.255 |
| Maximum length (μm) | |
| 1st day | 222.79 |
| 7th day | 164.84 |
| Minimum length (μm) | |
| 1st day | 190.18 |
| 7th day | 132.43 |
| Standart deviation | |
| 1st day | 10.97027077 |
| 7th day | 10.06698748 |
| Variance | |
| 1st day | 120.3468408 |
| 7th day | 101.3442368 |
| Shapiro-Wilk test (α = 0.05) | |
| 1st day | W: 0.9273 |
| p-value: 0.1371 | |
| 7th day | W: 0.9556 |
| p-value: 0.4607 | |
| Student’s t-test | |
| t: 17.241 | |
| p: 1.47.10-19 | |
| Critical t-value (p = 0.05): 2.0244 |
Discussion
The similarity of the morphological parameters between the specimens analyzed in the present study and the data presented in the description of L. trichoglossa ensures the correct taxonomic identification of the organisms. It should be noted that the characterization performed by Foissner (2003) occurred a few days after collection, whereas in the present work, the morphological study techniques were performed a few hours after collection. The temporal analyses on body length performed in the present work demonstrated a decrease in the average size of the organisms over time (Fig. 2), which would justify the slightly higher average values compared to the type population.
Regarding the occurrence of specimens with a greater number of caudal cilia, polymorphic life cycles are a common characteristic of the genus Tetrahymena. Lynn and Doerder (2012) mentioned the occurrence of changes in size, body shape, organization, morphology, and cilia of the oral apparatus of Tetrahymena species, due to environmental changes. The aforementioned authors highlighted the great genotypic and phenotypic plasticity found in the clade.
Phylogenetic reconstructions performed with both markers demonstrate the internal position of L. trichoglossa in the genus Tetrahymena, corroborating analyses based on the SSU marker presented by Strüder-Kypke et al. (2001), Bourland and Strüder-Kypke (2010), and Dunthorn et al. (2012). Several authors demonstrated the division of the genus Tetrahymena into 2 strongly molecularly supported clades, Australis and Borealis (Chantangsi, 2007; Chantangsi & Lynn, 2008; Lynn et al., 2018; Nanney, 1998; Strüder-Kypke et al., 2001). This division was observed in the inferred reconstructions by the SSU marker, performed in the present study, demonstrating the inclusion of L. trichoglossa into the Borealis clade. Doerder (2018) states that this clade has greater molecular diversity and more species than the Australis clade, thus being an important source of diversity for the genus. Lambornella trichoglossa, included in this clade, is the greatest example of phenotypic diversification, given its morphological and ontogenetic particularities demonstrated by Foissner (2003).
The phylogenetic analyses inferred by the COI gene demonstrated the inclusion of L. trichoglossa in a clade called coxset A4 by Chantangsi and Lynn (2008). The species T. corlissi and T. bergeri, members of this clade, share the parasitic lifestyle and have particular morphological characteristics (Hoffman et al., 1975; Imai et al., 2000; Strüder-Kypke et al., 2001). Foissner (2003) performed infection tests with L. trichoglossa and different mosquito species, noting that the ciliate does not develop a parasitic association. However, congeners L. stegomyiae and L. clarki are parasites of culicid dipterans, demonstrating that this way of life is predominant in coxiset A4, corroborating Chantangsi and Lynn (2008). Probably, parasitism also represents the ancestral condition of this clade, since Strüder-Kypke et al. (2001) demonstrated that histophagy is a recurrent phenomenon in the evolution of Tetrahymena.
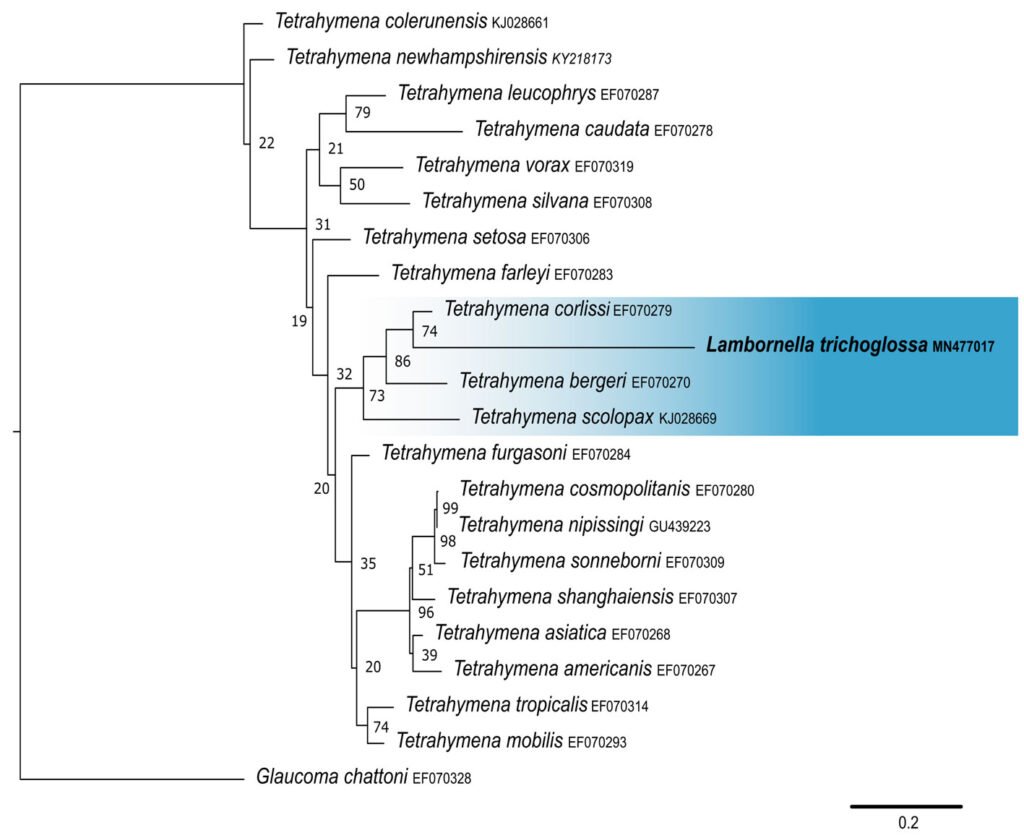
Figure 4. Maximum likelihood (ML) tree inferred from cytochrome oxidase subunit I (COI). Bold: sequence obtained in the present work. Numbers at nodes represent the bootstrap values of ML out of 500 replicates.
Table 3
Pairwise distance between SSU sequences of Lambornella trichoglossa and Tetrahymena spp.
| MN567691 | L. trichoglossa | ||||||||
| AJ810078 | L. trichoglossa* | 0.0029 | |||||||
| JQ723973 | Lambornella sp.** | 0.0053 | 0.0047 | ||||||
| U17356 | T. corlissi | 0.0059 | 0.0063 | 0.0077 | |||||
| AF364039 | T. bergeri | 0.0071 | 0.0092 | 0.0089 | 0.0046 | ||||
| EF070247 | T. furgasoni | 0.0130 | 0.0149 | 0.0149 | 0.0099 | 0.0092 | |||
| AF364040 | T. mobilis | 0.0130 | 0.0151 | 0.0149 | 0.0104 | 0.0092 | 0.0006 | ||
| KJ028502 | T. colerunensis | 0.0132 | 0.0139 | 0.0139 | 0.0087 | 0.0080 | 0.0051 | 0.0058 | |
| KJ028504 | T. scolopax | 0.0135 | 0.0156 | 0.0156 | 0.0094 | 0.0067 | 0.0135 | 0.0142 | 0.0125 |
Bold font: sequence obtained in the present study. * Sequence obtained from the type population (Dunthorn et al., 2012). ** Sequences obtained from samples collected in Jamaica (Dunthorn et al., 2012).
Table 4
Pairwise distance between COI sequences of Lambornella trichoglossa and Tetrahymena spp.
| MN477017 | L. trichoglossa | |||||||||
| EF070279 | T. corlissi | 0.1294 | ||||||||
| EF070270 | T. bergeri | 0.1450 | 0.0921 | |||||||
| EF070291 | T. malaccensis | 0.1536 | 0.1187 | 0.1320 | ||||||
| EF070268 | T. asiatica | 0.1557 | 0.1026 | 0.1113 | 0.1062 | |||||
| KY218158 | T. alphathermophila | 0.1713 | 0.1152 | 0.1403 | 0.1175 | 0.1281 | ||||
| EF070296 | T. paravorax | 0.1961 | 0.1400 | 0.1491 | 0.1413 | 0.1185 | 0.1407 | |||
| KJ028669 | T. scolopax | 0.1982 | 0.1364 | 0.1376 | 0.1461 | 0.1184 | 0.1534 | 0.1491 | ||
| EF070278 | T. caudata | 0.1857 | 0.1429 | 0.1492 | 0.1622 | 0.1331 | 0.1553 | 0.1735 | 0.1615 | |
| MF693881 | T. glochidiophila | 0.2187 | 0.1639 | 0.1867 | 0.1626 | 0.1631 | 0.1703 | 0.1500 | 0.1926 | 0.2051 |
Bold font: sequence obtained in the present study.
Traditionally recognized Tetrahymena species, described based on morphological or biochemical characteristics, and molecularly supported, may have high evolutionary difference values when comparing COI sequences. For example, T. caudata differs by over 15% from T. corlissi, T. termophila, and T. glochidiophila. Some species described by Doerder (2018) based mainly on molecular diversity differ by values close to 20%. Therefore, the 12.94% distance observed between the COI sequence of L. trichoglossa and T. corlissi does not represent a discrepancy concerning the molecular variability observed in Tetrahymena.
According to Chantangsi and Lynn (2008), the mitochondrial COI gene is effective in elucidating Tetrahymena’s recent phylogeny, while the SSU marker is more suited to assessing the deep phylogeny of the genus. The maintenance of topology in the reconstructions performed with both markers demonstrates that L. trichoglossa, T. corlissi, and T. bergeri share a long evolutionary history, and the differentiation of the species belonging to the genus Lambornella could be recent within the phylogeny of Tetrahymena. The phylogenetic position of L. trichoglossa also demonstrated by the COI gene, reinforces the proposals for synonymization of the genera Lambornella and Tetrahymena. Although Lambornella’s description precedes that of Tetrahymena, the principle of stability is appropriate to maintain the name of the latter taxon. In any case, it is necessary to establish an extended diagnosis for Tetrahymena, so that the species involved in new combinations are covered.
Finally, the molecular analysis of L. trichoglossa based on the COI marker supports the proposal for synonymization of the genera Lambornella and Tetrahymena. This systematic readjustment also points to the great molecular, morphological, evolutionary and ecological complexity of the genus Tetrahymena, which represents one of the greatest examples of evolutionary adaptation among eukaryotic organisms.
References
Bourland, W. A., & Strüder-Kypke, M. (2010). Agolohymena aspidocauda nov. gen., nov.spec., a histophagous freshwater tetrahymenid ciliate in the family Deltopylidae (Ciliophora, Hymenostomatia), from Idaho (northwest U.S.A.): morphology, ontogenesis and molecular phylogeny. European Journal of Protistology, 46, 221–242. https://doi.org/10.1016/j.ejop.2010.04.003
Buosi, P. R. B., Utz, L. R. P., Meira, B. R., Silva, B. T. S., Lansac-Tôha, F. M., Lansac-Tôha, F. A. et al. (2014). Rainfall influence on species composition of the ciliate community inhabiting bromeliad phytotelmata. Zoological Studies, 53, 32–44. https://doi.org/10.1186/s40555-014-0032-4
Buosi, P. R. B., Cabral, A. F., Utz, L. R. P., Vieira, L. C. G., & Velho, L. F. M. (2015). Effects of seasonality and dispersal on the ciliate community inhabiting bromeliad phytotelmata in riparian vegetation of a large tropical river. Journal of Eukaryotic Microbiology, 62, 737–749. https://doi.org/10.1111/jeu.12232
Chantangsi, C., & Lynn, D. H. (2008). Phylogenetic relationships within the genus Tetrahymena inferred from the cytochrome c oxidase subunit 1 and the small subunit ribosomal RNA genes. Molecular Phylogenetics and Evolution, 49, 979–87. https://doi.org/10.1016/j.ympev.2008.09.017
Chantangsi, C., Lynn, D. H., Brandl, M. T., Cole, J. C., Hetrick, N., & Ikonomi, P. (2007). Barcoding ciliates: a comprehensive study of 75 isolates of the genus Tetrahymena. International Journal of Systematic and Evolutionary Microbiology, 57, 2412–2423. https://doi.org/10.1099/ijs.0.64865-0
Corliss, J. O., & Coats, D. W. (1976). A new cuticular cyst-producing tetrahymenid ciliate, Lambornella clarki n. sp., and the current status of ciliatosis in culicine mosquitoes. Transactions of the American Microscopical Society, 95, 725–739. https://doi.org/10.2307/3225400
Corliss, J. O. (1960). Tetrahymena chironomi sp. nov., a ciliate from midge larvae, and the current status of facultative parasitism in the genus Tetrahymena. Parasitology, 50, 111–153. https://doi.org/10.1017/S0031182000025245
Doerder, F. P. (2018). Barcodes reveal 48 new species of Tetrahymena, Dexiostoma, and Glaucoma: phylogeny, ecology, and biogeography of new and established species. Journal of Eukaryotic Microbiology, 66, 182–208. https://doi.org/10.1111/jeu.12642
Dunthorn, M., Stoeck, T., Wolf, K., Breiner, H., & Foissner, W. (2012). Diversity and endemism of ciliates inhabiting Neotropical phytotelmata. Systematics and Biodiversity, 10, 195–205. https://doi.org/10.1080/14772000.2012.685195
Durán-Ramírez, C., García-Franco, J. G., Foissner, W., & Mayén-Estrada, R. (2015). Free-living ciliates from epiphytic tank bromeliads in Mexico. European Journal of Protistology, 51, 15–33. https://doi.org/10.1016/j.ejop.2014.09.002
Foissner, W. (2003). Morphology and ontogenesis of Lambornella trichoglossa nov. spec., a new tetrahymenid ciliate (Protozoa,Ciliophora) from Brazilian tank bromeliads (Bromeliaceae). European Journal of Protistology, 39, 63–82. https://doi.org/10.1078/0932-4739-00887
Foissner, W. (2014). An update of basic light and scanning electron microscopic methods for taxonomic studies of ciliated protozoa. International Journal of Systematic and Evolutionary Microbiology, 64, 271–292. https://doi.org/10.1099/ijs.0.057893-0
Foissner, W., Strüder-Kypke, M., van der Staay, G. W. M., van der Staay, S. M., & Hackstein, J. H. P. (2003). Endemic ciliates (Protozoa,Ciliophora) from tank bromeliads (Bromeliaceae): a combined morphological, molecular, and ecological study. European Journal of Protistology, 39, 365–372. https://doi.org/0.1078/0932-4739-00005
Hammer, Ø., Harper, D. A. T., & Ryan, P. D. (2001). PAST: Paleontological Statistics Software Package for Education and Data Analysis. Palaeontologia Electronica, 4, 1–9.
Hoffman, G. L., Lando, M., Camper, J. E., Coats, D. W., Stookey, J. L., & Burek, J. D. (1975). A disease of freshwater fishes caused by Tetrahymena corlissi Thompson, 1955, and a key for identification of holotrich ciliates of freshwater fishes. The Journal of Parasitolgy, 61, 217–223. https://doi.org/10.1111/j.1365-2761.2009.01062.x
Imai, S., Tsurimaki, S., Goto, E., Wakita, K., & Hatai, K. (2000). Tetrahymena infection in Guppy, Poecilia reticulata. Fish Pathology, 35, 67–72. https://doi.org/10.3147/jsfp.35.67
Katoh, K., Rozewicki, J., & Yamada, K. D. (2017). MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20, 1160–1166. https://doi.org/10.1093/bib/bbx108
Keilin, D. (1921). On a new ciliate: Lambornella stegomyiae n. g., n. sp., parasitic in the body-cavity of the larvae of Stegomyia scutellaris Walker (Diptera, Nematocera, Culicidae). Parasitology, 13, 216–224. https://doi.org/10.1017/s0031182000012488
Kimura, M. (1980). A simple method for estimating evolutionary rate of base substitutions through comparative studies
of nucleotide sequences. Journal of Molecular Evolution, 16, 111–120. https://doi.org/10.1007/BF01731581
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Molecular Biology and Evolution, 35, 1547–1549. https://doi.org/10.1093/molbev/msy096
Lynn, D. H., & Doerder, F. P. (2012). The life and times of Tetrahymena. In Collins, K. Methods in cell biology. Elsevier, 109, 9–27. https://doi.org/10.1016/B978-0-12-385
967-9.00002-5
Lynn, D. H., Doerder, F. P., Gillis, P. L., & Prosser, R. S. (2018). Tetrahymena glochidiophila n. sp., a new species of Tetrahymena (Ciliophora) that causes mortality to glochidia larvae of freshwater mussels (Bivalvia). Diseases of Aquatic Organisms, 127, 125–136. https://doi.org/10.3354/dao03188
Lynn, D. H. (2008). The ciliated taxa including families and genera. In D. H. Lynn (Ed.), The ciliated protozoa (pp. 339–440). Guelph, Canada: Springer.
Messing, J. (1983). New M13 vectors for cloning. Methods in Enzymology, 101, 20–78. https://doi.org/10.1016/0076-6879(83)01005-8
Modeo, L., Rosati, L., Andreoli, I., Simoni, G., Verni, F., & Petroni, G. (2006). Molecular systematics and ultrastructural characterization of a forgotten species: Chattonidium setense (Ciliophora, Heterotrichea). Proceedings of the Japan Academy, 82, 359–374. https://doi.org/10.2183/pjab.82.359
Nanney, D. L., Park, C., Preparata, R., & Simon, E. M. (1998). Comparison of sequence differences in a variable 23S rRNA domain among sets of cryptic species of ciliated protozoa. Journal of Eukaryotic Microbiology, 45, 91–100. https://doi.org/10.1111/j.1550-7408.1998.tb05075.x
Schrallhammer, M., Ferrantini, F., Vannini, C., Galati, S., Schweikert, M., Görtz, H. et al. (2013). Candidatus Megaira polyxenophila gen. nov., sp. nov.: Considerations on evolutionary history, host range and shift of early divergent Rickettsiae. Plos One, 8, e72581. https://doi.org/10.1371/journal.pone.0072581
Stamatakis, A. (2014). RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30, 1312–3. https://doi.org/10.1093/bioinformatics/btu033
Strüder-Kypke, M., Wright, H. D., Jerome, C. A, & Lynn, D. H. (2001). Parallel evolution of histophagy in ciliates of the genus Tetrahymena. Evolutionary Biology, 1, 1–5. https://doi.org/10.1186/1471-2148-1-5
Strüder-Kypke, M., & Lynn, D. H. (2010). Comparative analysis of the mitochondrial cytochrome c oxidase subunit I (COI) gene in ciliates (Alveolata, Ciliophora) and evaluation of its suitability as a biodiversity marker. Systematics and Biodiversity, 8, 131–148. https://doi.org/10.1080/14772000903507744
Tamura, K., Nei, M., & Kumar, S. (2004). Prospects for inferring very large phylogenies by using the neighbor-joining method. Proceedings of the National Academy of Sciences (USA), 101, 11030–11035. https://doi.org/10.1073/pnas.0404206101
Talavera, G., & Castresana, J. (2007). Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Systematic Biology, 56, 564–577. https://doi.org/10.1080/10635150701472164