Epicoccum zapotae: una especie nueva aislada de frutos de Manilkara zapota en el centro de México
Limni Silday Ramírez-Gallegos a, Laura Navarro-de la Fuente b, Ángel Trigos b, Irene Lagunes b, *
a Universidad Veracruzana, Centro de Investigación en Micología Aplicada, Doctorado en Micología aplicada, Calle Médicos 5, U.H. Del Bosque, 91010 Xalapa, Veracruz, Mexico
b Universidad Veracruzana, Centro de Investigación en Micología Aplicada, Calle Médicos 5, U.H. Del Bosque, 91010 Xalapa, Veracruz, Mexico
*Corresponding author: roslagunes@uv.mx (I. Lagunes)
Received: 07 October 2024; accepted: 03 November 2025
Abstract
The genus Epicoccum exhibits substantial intraspecific morphological and genetic diversity, which has made it difficult to correctly identify and delimit species on the basis of morphological characteristics alone. Against this background, this study was aimed to report the morphological and molecular identification of a new species belonging to the genus Epicoccum, isolated from sapodilla fruit (Manilkara zapota) in Veracruz, Mexico, named E. zapotae. The morphological characteristics and the phylogenetic analysis of the concatenated sequences of LSU, ITS, and β-TUB support the fact that the isolate is a new species of the genus Epicoccum. This discovery contributes to the knowledge of the diversity of Epicoccum species in tropical ecosystems.
Keywords: Didymellaceae; Epicoccum;Morphology; Phylogenetic; Sapodilla fruit
Resumen
El género Epicoccum presenta diversidad morfológica y genética intraespecífica considerable, lo cual ha dificultado la correcta identificación y delimitación de especies con base únicamente en características morfológicas. En este contexto, el objetivo de este estudio fue reportar la identificación morfológica y molecular de una especie nueva perteneciente al género Epicoccum, aislada del fruto chicozapote (Manilkara zapota) en Veracruz, México, denominada Epicoccum zapotae. Las características morfológicas y el análisis filogenético de las secuencias concatenadas de LSU, ITS y β-TUB respaldan el hecho de que el aislado es una nueva especie del género Epicoccum. Este descubrimiento contribuye al conocimiento de la diversidad de especies de Epicoccum en ecosistemas tropicales.
Palabras clave: Didymellaceae; Epicoccum;Morfología; Filogenética; Fruto del chicozapote
Introduction
The genus Epicoccum belongs to the Didymellaceae family. It exhibits substantial intraspecific morphological and genetic diversity. It can be found in the air, soil, water, and various parts of plants including flowers, branches, leaves, and bark (Chen et al., 2017; Fávaro et al., 2011; Lee et al., 2020; Li et al., 2022). More than 70 species of the genus Epicoccum have been described, of which 5 species (E. dendrobii, E. layuense, E. mezzettii, E. minitans, and E. nigrum) have demonstrated biocontrol activity against phytopathogens (Braga et al., 2018; Taguiam et al., 2021). Species of this genus may exhibit a saprophytic (Braga et al., 2018), phytopathogenic (E. nigrum and E. sorghinum) (Chen et al., 2017; Taguiam et al., 2021) or endophytic lifestyle, the latter having isolated compounds with antioxidant, anticancer and antimicrobial activities (Braga et al., 2018; Taguiam et al., 2021).
The genus Epicoccum was originally established by Link (1816) based on observations of sporodochia in dry plant stems, with the following diagnosis: compact globose stroma dotted with subglobose spores. Link (1816) described the type species E. nigrum, however, his diagnosis was unsatisfactory and generated confusion (Schol-Schwarz, 1959). Subsequently, many species belonging to this genus were described, based on the fungus characteristics in its natural habitat, and therefore, they were named of the plant species from which the specimen was isolated (Schol-Schwarz, 1959).
Schol-Schwarz (1959) conducted a thorough review of the Epicoccum genus, examining 70 specimens displaying a variety of hues, including red, yellow, or olive, alongside an additional 96 specimens from herbaria around the world. Despite this extensive examination, an incomplete diagnosis was noted with descriptions that were primarily based on the fungus’s original habitat characteristics and lacking the spore size measurements. For this reason, Schol-Schwarz (1959) considered the genus misclassified, concluding that color could not be used as a reliable taxonomic character, due to its susceptibility to various abiotic factors such as medium, light, pH, etc.; thus, no valid basis remained for splitting the genus into separate species, despite its wide range of conidial dimensions and cultural characters. Consequently, the entire genus was reduced to the single variable species E. nigrum (Kilpatrick & Chilvers, 1981).
Noting the lack of molecular evidence to support Schol-Schwarz´s classification, Wang and Guo (2004) performed the molecular identification of 45 E. nigrum isolates by amplifying the 5.8S gene and flanking internal transcribed spacer regions (ITS1 and ITS2) of the nuclear ribosomal DNA. They found that E. nigrum could comprise more than 1 species, ultimately concluding that conidial size and colony color should not be used as reliable taxonomic characteristics in the identification of E. nigrum. Later, Fávaro et al. (2011) acknowledged the need to reevaluate the classification of E. nigrum as a single variable species and many sequences deposited as E. nigrum in the GenBank database and fungal strains cultures many of which should be reclassified. Accurate species identification has long been challenging due to its heavy reliance on host plant morphology and association (Chen et al., 2015).
Therefore, taxonomic studies based on multilocus phylogeny using LSU, ITS, rpb2 and tub2 sequences in combination with morphological differences, allowed revising the classification within the Didymellaceae family, including the genus Epicoccum, providing a relatively robust phylogenetic basis for taxon delineation (Aveskamp et al., 2010; Braga et al.,2018; Chen et al., 2015; Chen et al., 2017). Currently, 18 Epicoccum species are accepted with their correct morphological and molecular identification (Braga et al., 2018). In this context, this study was aimed to report the morphological and molecular identification of a new species belonging to the genus Epicoccum.
Materials and methods
Sapodilla fruits were collected during May-June 2017 from 10-12 tall Manilkara zapota trees in one ~ 1 ha orchard close to the village of Apazapan, Veracruz, Mexico. Fruit samples were transported to the laboratory at the Centro de Investigación en Micología Aplicada (CIMA), Universidad Veracruzana. The samples were washed with running tap water to remove residual soil, afterwards the fruits were cut into small pieces (0.5 cm2), disinfected with 2% sodium hypochlorite for 30 seconds, and washed twice with sterile water. The disinfected samples were transferred onto Petri dishes containing Potato Dextrose Agar (PDA) medium (MCD LAB, San Jacinto Amilpas, Oax, MX) with 0.2 g/L chloramphenicol. The Petri dishes were incubated in dark conditions at 25 ± 2 °C for 2-3 days until mycelial growth was developed from the samples. Newly developed mycelia were immediately transferred to fresh PDA plates and incubated under the same conditions for fungal isolation. To ensure the purity of the fungal isolate, monosporic cultures were developed by cutting hyphal tips under a stereoscopic microscope (Leica EZ4), transferred to PDA plates and incubated under the same conditions established.
Culture characteristics were determined after 14 days of cultivation at 25 ± 2 °C on PDA. Colony diameters were measured after 7 days in incubation. The hyphae and reproductive structures produced on PDA were morphologically identified using an optical microscope at 100 × magnification. The mycelium was mounted on a microscope slide with a drop of lactophenol blue solution; the diameter of the conidia was measured (n = 100), means and standard errors (SE) were calculated, with extreme values shown in parentheses. The reproductive structures were identified using the taxonomic keys reported by Barnett and Hunter (1972). The holotype specimen (CM-CNRG 1003, dried agar plate culture) was deposited in the Microorganism Collection of the National Institute of Forestry, Agricultural and Livestock Research (INIFAP) in Mexico.
Genomic DNA was extracted from 14 days old mycelium grown on PDA according to Liu et al. (2000). The internal transcribed spacer region (ITS) of the nuclear ribosomal DNA (nrDNA) gene, the partial large subunit (LSU) nrDNA gene, and the β-tubulin (TUB) gene were amplified. PCR was performed using the primers listed in Table 1. DNA was amplified in a SureCycler 8800 thermal cycler with the same conditions for each gene, only the annealing temperature varied. The reaction mixture was incubated as follows: initial denaturation at 94 °C for 3 min, followed by 34 cycles at 94 °C for 1 min, the annealing temperatures are shown in Table 1, extension at 72 °C for 1 min, followed by a final extension at 72 °C for 10 min. The amplifications were purified using the Wizard® SV Gel and PCR System Clean-Up kit and sent to Labsergen Langebio (Cinvestav, Irapuato, Gto., Mexico) for sequencing on an AB3770 capillary sequencer.
Consensus sequences were created with BioEdit software (Hall, 1999) and were compared in the GenBank nucleotide sequence database (Benson et al., 2017) using BLAST search software (Zhang et al., 2000) to confirm the genus and percentage of identity. Closely related species sequences and our newly obtained sequences (Table 2) were incorporated into sequence datasets independently for each molecular marker using PhyDE v.0.9971 Phylogenetic Development (Müller et al.,2010). Each dataset was independently aligned using the MAFFT online service (Katoh et al., 2019). Inconsistencies were manually adjusted using the MESQUITE 3.61 phylogenetic data editor (Maddison & Maddison, 2019), the same program with which the concatenated ITS + LSU + TUB sequences dataset were integrated. The GTR + G nucleotide substitution model was selected using the jModelTest v.2.1.4 program (Darriba et al., 2012) according to the Akaike Information Criterion (AIC). The concatenated aligned dataset was analyzed using Maximum Likelihood (ML) and Bayesian Inference (BI). The ML analyses were conducted with RAxMLGUI2.0 (Edler et al., 2020) using 1,000 bootstrap replicates, the robustness of the analyses was evaluated by bootstrap support (BS) values. Bayesian Inference analyses were performed on MrBayes v.3.2.1 software, using the substitution model selected for each matrix by jModelTest, based on the Akaike Information Criterion (AIC) (Table 3), with 4 Markov Chains Monte Carlo (MCMC) and 1 million generations (Ronquist et al., 2012). The ITS, LSU and TUB sequences generated in this study were deposited in the GenBank database under the accession numbers provided in the taxonomic section.
The Genealogical Concordance Phylogenetic Species Recognition (GCPSR) model (Taylor et al., 2000) was used to delimit the fungal species and analyze phylogenetic relationships among closely related taxa. Recombination levels within these phylogenetically proximate species were assessed through the calculation of the pairwise homoplasy index (PHI) using the SplitsTree4 software (Huson, 1998; Huson & Bryant, 2006). A concatenated dataset including ITS, LSU, and TUB was used for the analyses. Splits graphs were generated in SplitsTree4 utilizing both the Log-Det transformation and splits decomposition options, facilitating the visualization of the phylogenetic relationships among the species. Interpretation of a pairwise homoplasy index value falling below the 0.05 significance threshold (Φw < 0.05) was taken to signify the presence of recombination within the analyzed dataset.
Table 1
Primers and annealing temperatures used in the PCR amplifications step.
| Locus | Primer | Primer DNA sequences (5´–3´) | Annealing T (°C) / Time (s) | Reference |
| ITS | ITS1F | CTTGGTCATTTAGAGGAAGTAA | 53/45 | Gardes & Bruns, 1993; White et al., 1990 |
| ITS4 | TCCTCCGCTTATTGATATGC | |||
| LSU | LR0R | ACCCGCTGAACTTAAGC | 55/60 | Vilgalys & Hester, 1990 |
| LR7 | TACTACCACCAAGATCT | |||
| TUB | TUB1 | AACATGCGTGAGATTGTAAGT | 57/60 | Woudenberg et al., 2009 |
| TUB22 | TCTGGATGTTGTTGGGAATCC |
Table 2
Species and GenBank accession number of sequences used for the construction of the phylogenetic tree.
| Species | Voucher | Country | GenBank accession number | ||
| ITS | LSU | TUB | |||
| E. brahmansense | CBS:990.95 | Papua New Guinea | MN973514 | MN943720 | MT005614 |
| E. brahmansense | CBS 985.95 | Papua New Guinea | MN973513 | MN943719 | MT005613 |
| E. camelliae | LC:4862 | China | KY742092 | KY742246 | KY742334 |
| E. dendrobii | LC:8145 T | China | KY742093 | KY742247 | KY742335 |
| E. draconis | CBS 186.83 | Rwanda | GU237795 | GU238070 | GU237607 |
| E. duchesneae | CBS 218.81 | India | MN972935 | MN973322 | MN983950 |
| E. duchesneae | LC:5139 T | China | KY742095 | KY742249 | KY742337 |
| E. hordei | LC:8149 | Australia | KY742098 | KY742252 | KY742340 |
| E. hordei | LC:8148 T | Australia | KY742097 | KY742251 | KY742339 |
| E. huancayense | CBS:105.80 | Peru | MH861244 | MH873016 | GU237615 |
| E. huancayense | CBS 390.93 | Peru | GU237857 | GU238085 | GU237616 |
| E. layuense | LC:8155 T | China | KY742107 | KY742261 | KY742349 |
| E. mezzettii | CBS 173.38 | Italy | MN973496 | MN943701 | MT005596 |
| E. mezzettii | CBS 238.59 | No data | MN973494 | MN943699 | MT005594 |
| E. multiceps | CBS:119734 | Brazil | MN973512 | MN943718 | MT005612 |
| E. nigrum | CBS 173.73 | USA | FJ426996 | GU237975 | FJ427107 |
| E. nigrum | CBS 125.82 | Netherlands | FJ426995 | GU237974 | FJ427106 |
| E. oryzae | CBS:174.34 | Japan | MN973500 | MN943705 | MT005600 |
| E. oryzae | CBS:173.34 | Japan | MN973499 | MN943704 | MT005599 |
| E. ovisporum | CBS 180.80 T | No data | NR158228 | NG069492 | FJ427174 |
| E. plurivorum | CBS 558.81 | New Zealand | GU237888 | GU238132 | GU237647 |
| E. poaceicola | CBS:987.95 | Papua New Guinea | MN972955 | MN973343 | MN983969 |
| E. poae | LC:8161 | USA | KY742114 | KY742268 | KY742356 |
| E. poae | LC:8160 T | USA | KY742113 | KY742267 | KY742355 |
| E. proteae | CBS:114179 T | South Africa | MH862956 | MH874519 | LT623230 |
| E. tobaicum | JCK-CSHF10 | South Korea | MW368670 | MW368670 | MW392085 |
| E. variabile | CBS:119733 | Brazil | MN973501 | MN943706 | MT005601 |
| E. viticis | CGMCC 3.18344 T | China | NR158267 | NG069447 | KY742360 |
| E. zapotae | CM-CNRG 1003 | Mexico | OQ333010 | OQ333009 | OQ330858 |
| D. americana | CBS 185.85 | USA | FJ426972 | GU237990 | FJ427088 |
T Indicates type strains
Table 3
The nucleotide substitution model for each matrix obtained by jModelTest based on the Akaike Information Criterion.
| Locus | Length | Substitution model | MrBayes parameters (Nst-rates) |
| ITS | 455 pb | TIM2 + I + G | 6-invgamma |
| LSU | 765 pb | TrN + I + G | 6-invgamma |
| TUB | 330 pb | TIM3 + G | 6-gamma |
Description
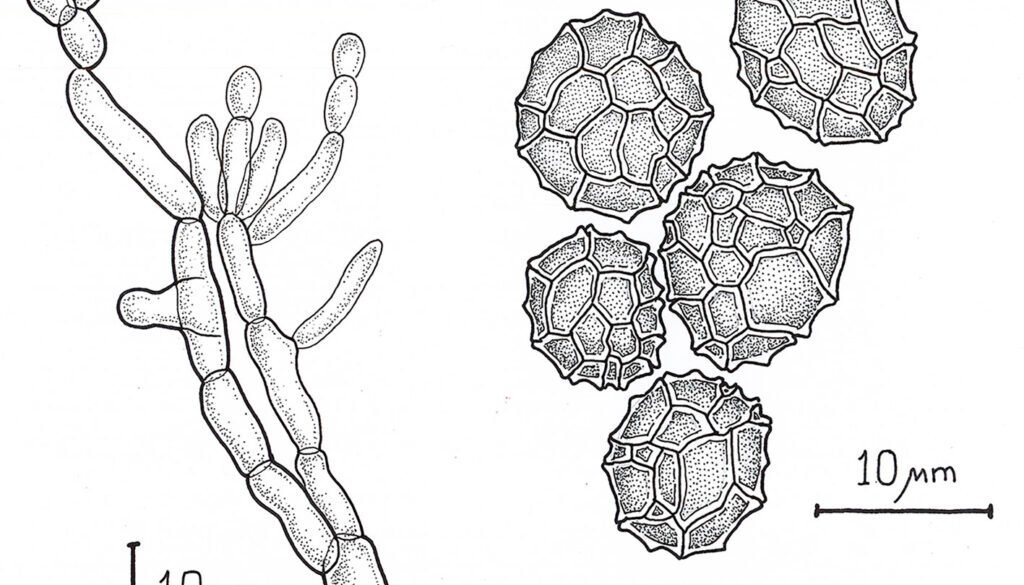
Epicoccum zapotae L. Navarro & L. Ramírez sp. nov. (Fig. 1)
Diagnosis: dark sporodochia, conidia dark pigmented, globose or obovoid with radial separation and a verrucous outer surface. Compared to other species in the genus, colonies growth on PDA exhibits a distinctive pink pigmentation.
Sexual morph: not observed.
Asexual morph: Sporodochia aredark. Conidiophores are compact or loose, and light brown. Conidia dark pigmented, globose or obovoid with radial septation and a verrucous outer surface, with an average size 10.48 ± 1.64 µm (8.5-13.4 µm) (Fig. 2).
Culture characteristics: Colonies on PDA measured 33 ± 2 mm in diameter after 7 days, furrowed and circular with filamentous margin and flattened aerial mycelia. The central region of the colonies appears light pink, surrounded by concentric rings of intense pink coloration.
This strain was deposited at Colección de Microorganismos del Centro Nacional de Recursos Genéticos (CNRG), Instituto Nacional de Investigaciones Forestales, Agrícolas y Pecuarias (INIFAP), Tepatitlán de Morelos, Jalisco, Mexico, under accession number CM-CNRG 1003.
Taxonomic summary
Mycobank number: MB 853124
GeneBank: OQ333010 (ITS), OQ333009 (LSU), OQ330858 (B-TUB)
Type: Mexico, Veracruz State: Apazapan (19°20’17.16” N, 96°43’54.12” W, 300 m), isolated from the fruit of M. zapota, June 2017, Holotype, CM-CNRG 1003.
Etymology: zapotae, referring to the host plant species (Manilkara zapota) from which it was isolated.
Remarks
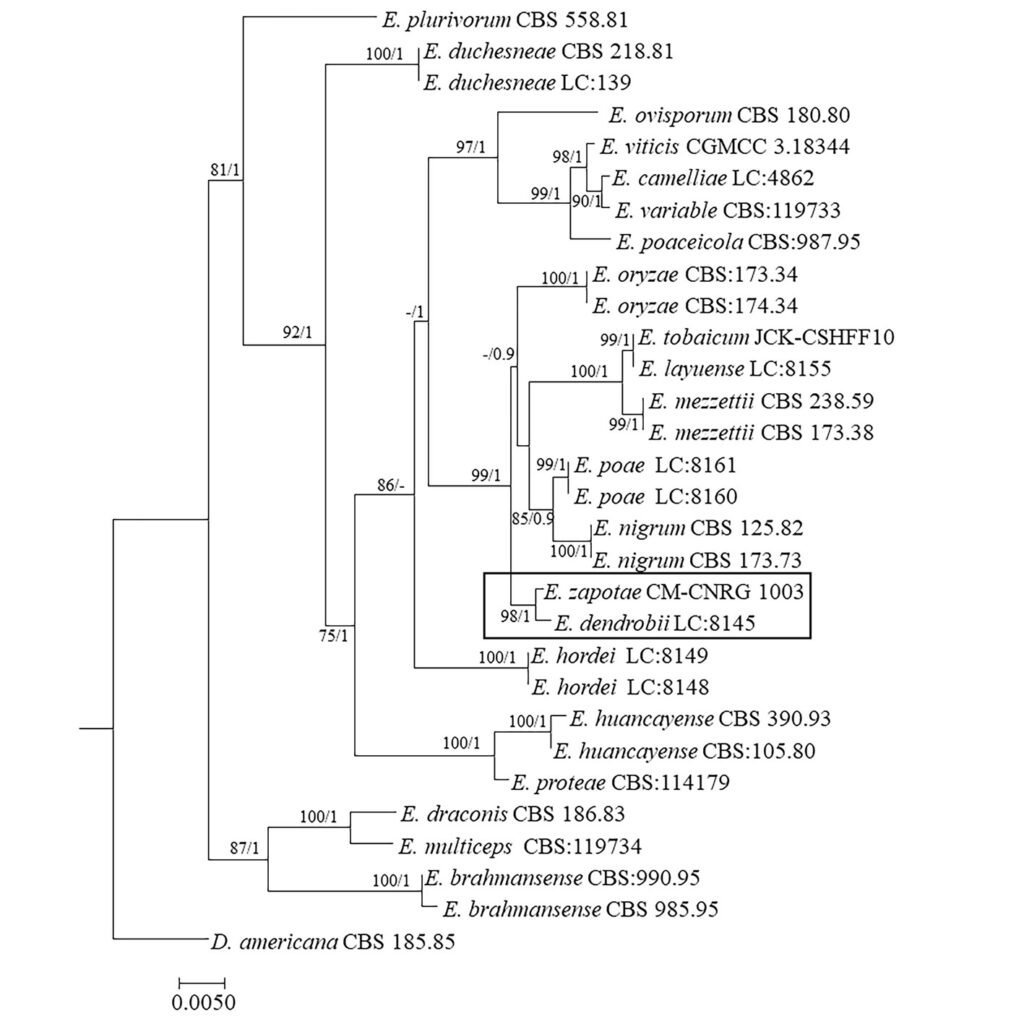
Phylogenetic tree constructed from concatenated ITS + LSU + TUB sequences of 30 different species, with a final length of 1,550 characters including gaps. Sequences of Didymella americana CBS 185.85 were selected as outgroup (Table 2, Fig. 3). Branches of the phylogenetic tree are labeled with their respective bootstrap values (BS) and the Bayesian posterior probabilities (BPP). Our consensus phylogenetic tree generated from the ML analysis with BS/BPP values robustly supports (BS = 98 / BPP = 1) the assignment of our isolate as an independent species within the Epicoccum genus, positioned as a sister taxon to E. dendrobii (Fig. 3).
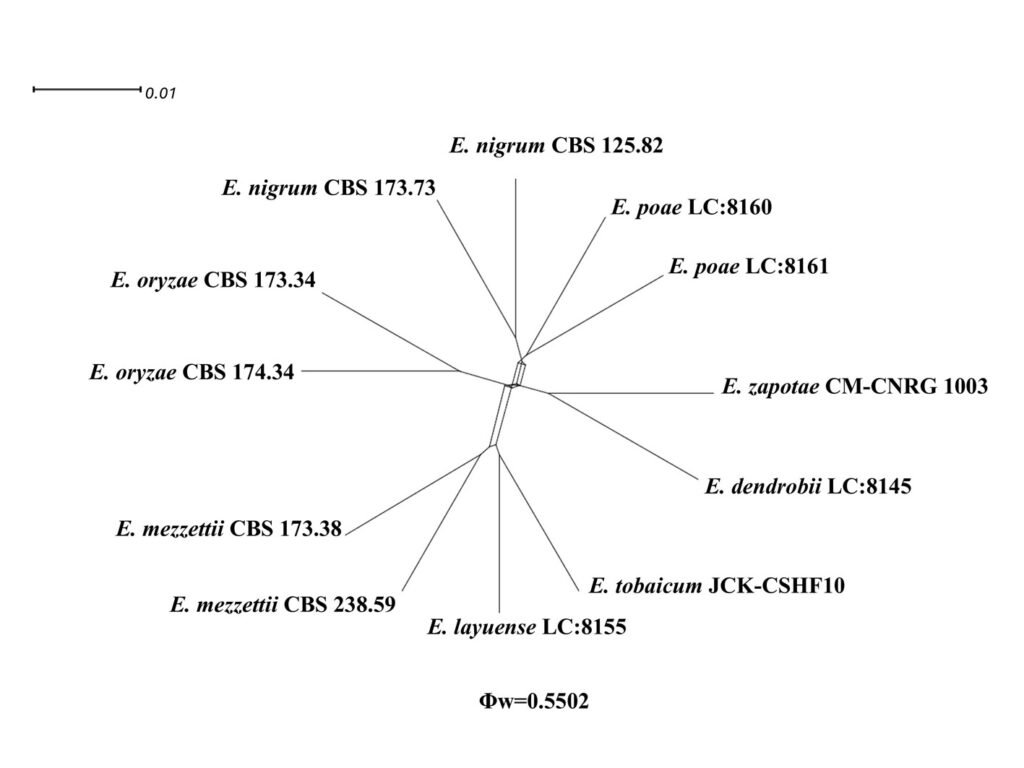
Application of the GCPSR analysis to our dataset yielded a PHI value of 0.5502 (Fig. 4). This outcome suggests that there is no substantial evidence of recombination occurring between E. zapotae and its related taxa. These results indicate that the E. zapotae isolate is distinctly differentiated from E.dendrobii and there is no indication of recombination events between them.
The robustness of BS/BPP values 98/1 supports the placement of E. zapotae as a sister species to E. dendrobii. This relationship forms a distinct clade, positioned adjacent to E. layuense, E. mezzettii, E. nigrum, E. oryzae, E. poae, and E. tobaicum.
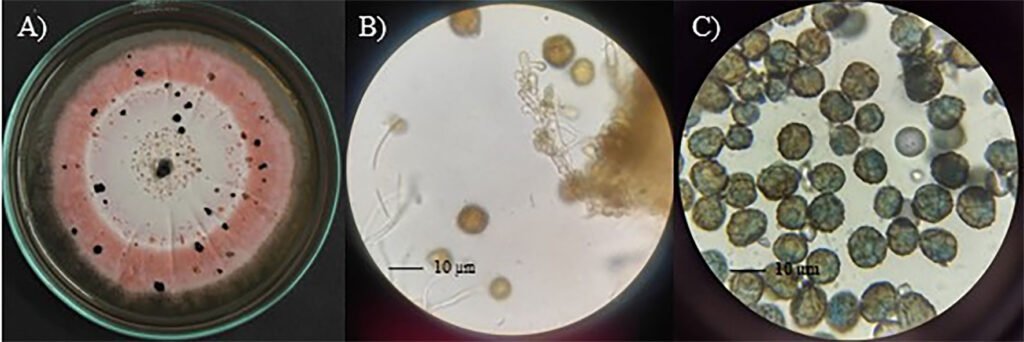
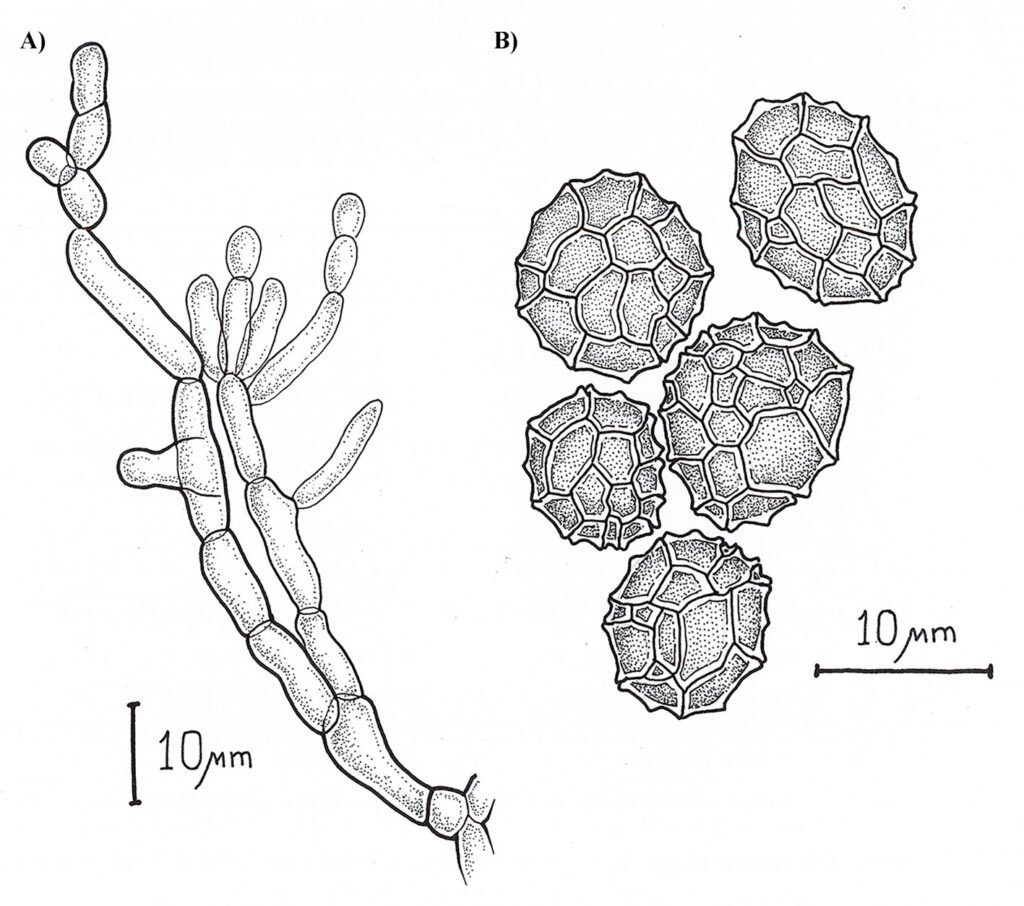

Epicoccum dendrobii exhibits the closest genetic proximity to E. zapotae. Although E. zapotae shares general morphological traits with other Epicoccum species, distinct features support the classification of E. zapotae as a separate species. Conidia of E. dendrobii are multicellular-phragmosporous, verrucose, subglobose-pyriform, brown, with a basal cell, 11-19 µm diam (Chen et al., 2017). In contrast, conidia of E. zapotae are smaller, ranging from 8.84-12.12 µm diam. Additionally, the radial growth rate of E. dendrobii on PDA at 25 °C reaches 34-38 mm diam after 7 days. There is also a significant difference in culture characteristics on PDA between the 2 species, colonies of E. dendrobii display a regular margin, with flat aerial mycelia felty to floccose in texture, white to buff in color, becoming olivaceous near the center (Chen et al., 2017). In contrast, colonies of E. zapotae show slower growth on PDA (31-35 mm) for 7 days and their colonies are distinguished by their intense pink color (Table 4).
Table 4
Comparative morphological characteristics between E. dendrobii and E. zapotae. Data for E. dendrobii adapted from Chen et al. (2017).
| Characteristic | E. dendrobii | E. zapotae |
| Conidia | Multicellular-phragmosporous, verrucose, subglobose-pyriform, brown | Dark pigmented, globose to obovoid with radial separation and a verrucous outer surface |
| Conidial diameter | 11-19 µm | 8.84-12.12 µm |
| Colony margin (PDA) | Regular | Furrowed and circular |
| Aerial mycelium (PDA) | Felty to floccose | Flattened |
| Colony color (PDA) | Beige | Pink |
| Colony diameter (7 days) | 34-38 mm | 31-35 mm |
Epicoccum dendrobii has been reported to cause black leaf spots on Dendrobium fimbriatum (Chen et al., 2017). However, E. zapotae was isolated from sapodilla fruits without visible disease symptoms; therefore, it was probably plant-associated as an endophytic fungus, that can grow asymptomatically within plant tissues (Wen et al., 2022). The ability of endophytic fungi to produce bioactive compounds with biomedical applications has been widely explored (Hashem et al., 2023). In particular, from Epicoccum sp. known as an endophyte of Theobroma cacao, the bioactive compounds epicolactone, epicoccolide A and epicoccolide B have been isolated (Talontsi et al., 2013). Therefore, E. zapotae represents an opportunity for mycochemical studies aimed at exploring the biomedical potential of the bioactive compounds it produced.
Our phylogenetic and morphological analysis, as well as the results from the GCPSR analysis confirm that our isolate, despite its greater genetic proximity to E. dendrobii, presents distinct morphological differences ranging from the size and shape of the conidia to their growth on PDA. This demonstrates that our isolate represents a new species of the genus Epicoccum.
Our comprehensive morphological and molecular analyses have led us to conclude that fungal isolate recovered from the M. zapota fruit, is a new species within the genus Epicoccum which we report as E. zapotae. This discovery represents the first report of E. zapotae isolated from the sapodilla fruits in Veracruz, Mexico, and provides an opportunity for bioprospecting studies aimed at exploring its pharmaceutical potential.
Acknowledgements
This research was supported by Secretaría de Ciencia, Humanidades, Tecnología e Innovación (Secihti), formerly Conahcyt (FORDECYT PRONACES CF/304020 project) and the Universidad Veracruzana (UV-CA-354). L.S.R-G thanks Secihti for the predoctoral grant 802758.
References
Aveskamp, M. M., de Gruyter, J., Woudenberg, J. H. C., Verkley, G. J. M., & Crous, P. W. (2010). Highlights of the Didymellaceae: a polyphasic approach to characterize Phoma and related pleosporalean genera. Studies in Mycology, 65, 1–60. https://doi.org/10.3114/sim.2010.65.01
Barnett, H. L., & Hunter, B. B. (1972). Illustrated genera of imperfect fungi. 3rd Ed. Minneapolis: Burgess Publishing Company.
Benson, D. A., Cavanaugh, M., Clark, K., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. et al. (2017). GenBank. Nucleic Acids Research, 45, D37–D42. https://doi.org/10.1093/nar/gkw1070
Braga, R., Padilla, G., & Araújo, W. L. (2018). The biotechnological potential of Epicoccum spp.: diversity of secondary metabolites. Critical Reviews in Microbiology, 44, 759–778. https://doi.org/10.1080/1040841X.2018.1514364
Chen, Q., Jiang, J. R., Zhang, G. Z., Cai, L., & Crous, P. W. (2015). Resolving the Phoma enigma. Studies in Mycology, 82, 137–217. https://doi.org/10.1016/j.simyco.2015.10.003
Chen, Q., Hou, L. W., Duan, W. J., Crous, P. W., & Cai, L. (2017). Didymellaceae revisited. Studies in Mycology, 87, 105–159. https://doi.org/10.1016/j.simyco.2017.06.002
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2012). JModelTest 2: More models, new heuristics and parallel computing. Nature Methods, 9, 772. https://doi.org/10.1038/nmeth.2109
Edler, D., Klein, J., Antonelli, A., & Silvestro, D. (2020). raxmlGUI 2.0: a graphical interface and toolkit for phylogenetic analyses using RAxML. Methods in Ecology and Evolution, 12, 373–377. https://doi.org/10.1111/2041-210X.13512
Fávaro, L. C., de Melo, F. L., Aguilar-Vildoso, C. I., & Araújo, W. L. (2011). Polyphasic analysis of intraspecific diversity in Epicoccum nigrum warrants reclassification into separate species. Plos One, 6, e14828. https://doi.org/10.1371/journal.pone.0014828
Gardes, M., & Bruns, T. D. (1993). ITS primers with enhanced specificity for basidiomycetes-application to the identifi-
cation of mycorrhizae and rusts. Molecular Ecology, 2,113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acid Symposium Series, 41, 95-98.
Hashem, A. H., Attia, M. S., Kandil, E. K., Fwazi, M. M., Abdelrahman, A. S., Khader, M. S. et al. (2023). Bioactive compounds and biomedical application of endophytic fungi: a recent review. Microbial Cell Factories, 22, 107. https://doi.org/10.1186/s12934-023-02118-x
Huson, D. H. (1998). SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics, 14, 68–73. https://doi.org/10.1093/bioinformatics/14.1.68
Huson, D. H., & Bryant, D. (2006). Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution, 23, 254–267. https://doi.org/10.1093/molbev/msj030
Katoh, K., Rozewicki, J., & Yamada, K. D. (2019). MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20, 1160–1166. https://doi.org/10.1093/bib/bbx108
Kilpatrick, J. A., & Chilvers, G. A. (1981). Variation in a natural population of Epicoccum purpurascens. Transactions of the British Mycological Society, 77, 497–508. https://doi.org/10.1016/S0007-1536(81)80097-6
Lee, A. J., Cadelis, M. M., Kim, S. H., Swift, S., Copp, B. R., & Villas-Boas, S. G. (2020). Epipyrone A, a broad-spectrum antifungal compound produced by Epicoccum nigrum ICMP 19927. Molecules, 25, 5997. https://doi.org/10.3390/molecules25245997
Li, T., Im, J., & Lee, J. (2022). Genetic diversity of Epicoccum nigrum and its effects on Fusarium graminearum. Mycobiology, 50, 457–466. https://doi:10.1080/12298093.2022.2148394
Link, H. F. (1816). Observationes in ordines plantarum naturales III. Magazin der Gesellschaft Naturforschender Freunde Berlin, 7, 37–38.
Liu, D., Coloe, S., Baird, R., & Pedersen, J. (2000). Rapid mini-preparation of fungal DNA for PCR. Journal of Clinical Microbiology, 38, 471. https://doi:10.1128/jcm.38.1.471-471.2000
Maddison, W., & Maddison, D. (2019). Mesquite: a modular system for evolutionary analysis. Version 3.61. https://www.mesquiteproject.org
Müller, J., Müller, K., Neinhuis, C., & Quandt, D. (2010). PhyDE Phylogenetic data editor, version 0.9971. Program distributed by the authors. http://www.phyde.de
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Höhna, S. et al. (2012). Mrbayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61, 539–542. https://doi.org/10.1093/sysbio/sys029
Schol-Schwarz, M. B. (1959). The genus Epicoccum link. Transactions of the British Mycological Society, 42, 149–173. https://doi.org/10.1016/S0007-1536(59)80024-3
Taguiam, J. D., Evallo, E., & Balendres, M. A. (2021). Epicoccum species: ubiquitous plant pathogens and effective biological control agents. European Journal of Plant Pathology, 159, 713–725. https://doi.org/10.1007/s10658-021-02207-w
Talontsi, F. M., Dittrich, B., Schüffler, A., Sun, H., & Laatsch, H. (2013) Epicoccolides: antimicrobial and antifungal polyketides from an endophytic fungus Epicoccum sp. associated with Theobroma cacao. European Journal of Organic Chemistry, 2013, 3174–3180. https://doi.org/10.1002/ejoc.201300146
Taylor, J. W., Jacobson, D. J., Kroken, S., Kasuga, T., Geiser, D. M., Hibbett, D. S. et al. (2000). Phylogenetic species recognition and species concepts in fungi. Fungal Genetics and Biology, 31, 21–32. https://doi.org/10.1006/fgbi.2000.1228
Vilgalys, R., & Hester, M. (1990). Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. Journal of Bacteriology, 172, 4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
Wang, Y., & Guo, L. D. (2004). Morphological and molecular identification of an endophytic fungus Epicoccum
nigrum. Mycosystema, 23, 474–479. https://doi:10.3329/bipt.v29i2.63534
Wen, J., Okyere, S. K., Wang, S., Wang, J., Xie, L., Ran, Y. et al. (2022). Endophytic fungi: an effective alternative source of plant-derived bioactive compounds for pharmacological studies. Journal of Fungi, 8, 205. https://doi.org/10.3390/jof8020205
White, T. J., Bruns, T. D., Lee, S. B., & Taylor, J. W. (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, & T. J. White (Eds.), PCR Protocols: a guide to methods and applications (pp. 315–322). New York: Academic Press. http://dx.doi.org/10.1016/B978-0-12-372180-8.50042-1
Woudenberg, J. H. C., Aveskamp, M. M., de Gruyter, J., Spiers, A. G., & Crous, P. W. (2009). Multiple Didymella teleomorphs are linked to the Phoma clematidina morphotype. Persoonia: Molecular Phylogeny and Evolution of Fungi, 22, 56–62. https://doi.org/10.3767/003158509X427808
Zhang, Z., Schwartz, S., Wagner, L., & Miller, W. (2000). A greedy algorithm for aligning DNA sequences. Journal of Computational Biology, 7, 203–214. https://doi.org/10.1089/10665270050081478